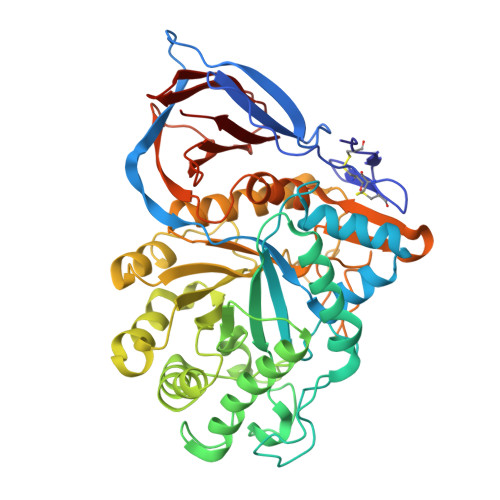
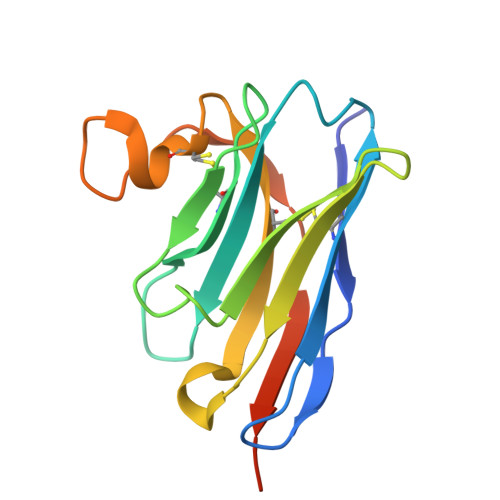
Developing nanobodies as allosteric molecular chaperones of glucocerebrosidase function.
Dal Maso, T., Sinisgalli, C., Zilio, G., Franzin, E., Tessari, I., Pardon, E., Steyaert, J., Ballet, S., Greggio, E., Versees, W., Plotegher, N.(2025) Nat Commun 16: 4890-4890
- PubMed: 40425544 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-60134-4
- Primary Citation Related Structures:
9ENA - PubMed Abstract:
The enzyme glucocerebrosidase (GCase) catalyses the hydrolysis of glucosylceramide to glucose and ceramide within lysosomes. Homozygous or compound heterozygous mutations in the GCase-encoding GBA1 gene cause the lysosomal storage disorder Gaucher disease, while heterozygous and homozygous mutations are the most frequent genetic risk factor for Parkinson's disease. These mutations commonly affect GCase stability, trafficking or activity. Here, we report the development and characterization of nanobodies (Nbs) targeting and acting as molecular chaperones for GCase. We identify several Nb families that bind with nanomolar affinity to GCase. Based on biochemical characterization, we group the Nbs in two classes: Nbs that improve the activity of the enzyme and Nbs that increase GCase stability in vitro. A selection of the most promising Nbs is shown to improve GCase function in cell models and positively impact the activity of the N370S mutant GCase. These results lay the foundation for the development of new therapeutic routes.
- VIB-VUB Center for Structural Biology, VIB, Brussels, Belgium.
Organizational Affiliation: