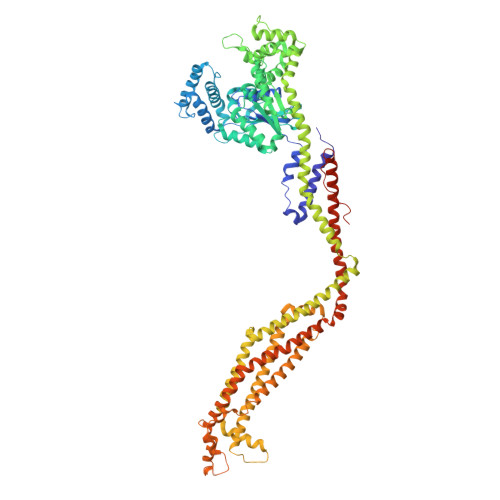
Structural basis for GTPase activity and conformational changes of the bacterial dynamin-like protein SynDLP.
Junglas, B., Gewehr, L., Mernberger, L., Schonnenbeck, P., Jilly, R., Hellmann, N., Schneider, D., Sachse, C.(2024) Cell Rep 43: 114657-114657
- PubMed: 39207903 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.114657
- Primary Citation Related Structures:
9EM7, 9EM8, 9EM9 - PubMed Abstract:
SynDLP, a dynamin-like protein (DLP) encoded in the cyanobacterium Synechocystis sp. PCC 6803, has recently been identified to be structurally highly similar to eukaryotic dynamins. To elucidate structural changes during guanosine triphosphate (GTP) hydrolysis, we solved the cryoelectron microscopy (cryo-EM) structures of oligomeric full-length SynDLP after addition of guanosine diphosphate (GDP) at 4.1 Å and GTP at 3.6-Å resolution as well as a GMPPNP-bound dimer structure of a minimal G-domain construct of SynDLP at 3.8-Å resolution. In comparison with what has been seen in the previously resolved apo structure, we found that the G-domain is tilted upward relative to the stalk upon GTP hydrolysis and that the G-domain dimerizes via an additional extended dimerization domain not present in canonical G-domains. When incubated with lipid vesicles, we observed formation of irregular tubular SynDLP assemblies that interact with negatively charged lipids. Here, we provide the structural framework of a series of different functional SynDLP assembly states during GTP turnover.
- Ernst Ruska-Center for Microscopy and Spectroscopy with Electrons (ER-C-3): Structural Biology, Forschungszentrum Jülich, 52425 Jülich, Germany.
Organizational Affiliation: