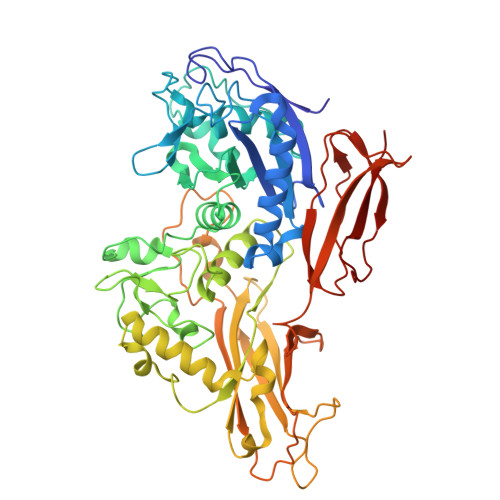
Structure of the Thomasclavelia ramosa immunoglobulin A protease reveals a modular and minimizable architecture distinct from other immunoglobulin A proteases.
Tran, N., Frenette, A., Holyoak, T.(2025) Proc Natl Acad Sci U S A 122: e2503549122-e2503549122
- PubMed: 40854123 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2503549122
- Primary Citation Related Structures:
9EKK, 9EKM, 9EKN - PubMed Abstract:
Immunoglobulin A proteases (IgAPs) are a diverse group of enzymes secreted from bacteria that inhabit human mucosal tissues. These enzymes have convergently evolved to cleave human immunoglobulin A as a means of modulating and evading host immunity. Only two of three known IgAP families have been biochemically characterized beyond their initial discovery. Here, we show using solution-scattering, steady-state kinetic, and crystallographic approaches that the protease from Thomasclavelia ramosa , representing the uncharacterized third family, has a truly modular and minimizable protein architecture. This analysis also revealed a unique metal-associated domain that likely functions as a molecular spacer and generated a working hypothesis detailing the structural mechanism behind the enzyme's high substrate specificity. Our work provides an in-depth biochemical account of this IgAP family, paving the way for advancing clinically relevant IgAP-related research and our understanding of IgAPs as a whole.
- Department of Biology, Faculty of Science, University of Waterloo, Waterloo, ON N2L 3G1, Canada.
Organizational Affiliation: