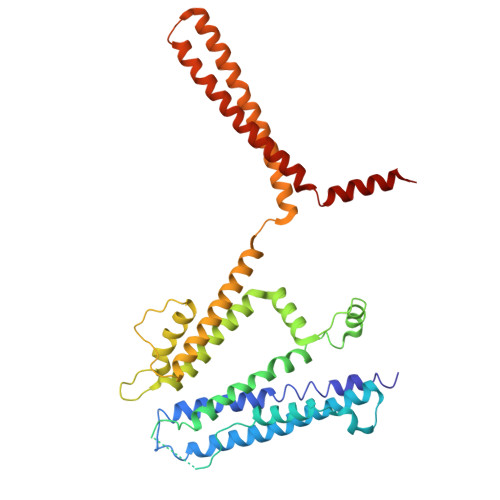
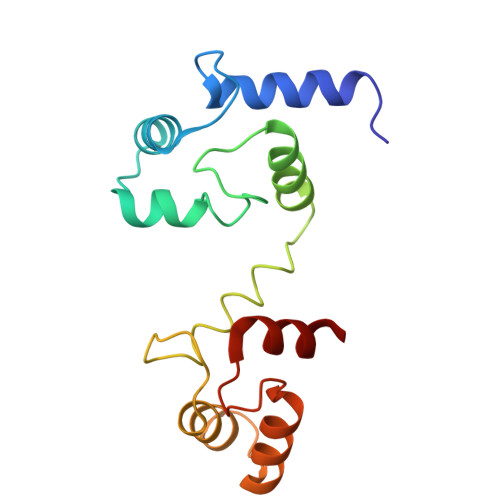
Design and structural basis of selective 1,4-dihydropyridine inhibitors of the calcium-activated potassium channel K Ca 3.1.
Ong, S.T., Nam, Y.W., Nasburg, J.A., Ramanishka, A., Ng, X.R., Zhuang, Z., Goay, S.S.M., Nguyen, H.M., Singh, L., Singh, V., Rivera, A., Eyster, M.E., Xu, Y., Alper, S.L., Wulff, H., Zhang, M., Chandy, K.G.(2025) Proc Natl Acad Sci U S A 122: e2425494122-e2425494122
- PubMed: 40294255 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2425494122
- Primary Citation Related Structures:
9ED1 - PubMed Abstract:
The 1,4-dihydropyridines, drugs with well-established bioavailability and toxicity profiles, have proven efficacy in treating human hypertension, peripheral vascular disorders, and coronary artery disease. Every 1,4-dihydropyridine in clinical use blocks L-type voltage-gated calcium channels. We now report our development, using selective optimization of a side activity (SOSA), of a class of 1,4-dihydropyridines that selectively and potently inhibit the intermediate-conductance calcium-activated K + channel K Ca 3.1, a validated therapeutic target for diseases affecting many organ systems. One of these 1,4-dihydropyridines, DHP-103, blocked K Ca 3.1 with an IC 50 of 6 nM and exhibited exquisite selectivity over calcium channels and a panel of >100 additional molecular targets. Using high-resolution structure determination by cryogenic electron microscopy together with mutagenesis and electrophysiology, we delineated the drug binding pocket for DHP-103 within the water-filled central cavity of the K Ca 3.1 channel pore, where bound drug directly impedes ion permeation. DHP-103 inhibited gain-of-function mutant K Ca 3.1 channels that cause hereditary xerocytosis, suggesting its potential use as a therapeutic for this hemolytic anemia. In a rat model of acute ischemic stroke, the second leading cause of death worldwide, DHP-103 administered 12 h postischemic insult in proof-of-concept studies reduced infarct volume, improved balance beam performance (measure of proprioception) and decreased numbers of activated microglia in infarcted areas. K Ca 3.1-selective 1,4-dihydropyridines hold promise for the many diseases for which K Ca 3.1 has been experimentally confirmed as a therapeutic target.
- Lee Kong Chian School of Medicine-Innovative CRO Explorer Collaborative Platform, Lee Kong Chian School of Medicine, Nanyang Technological University, Singapore 636921, Singapore.
Organizational Affiliation: