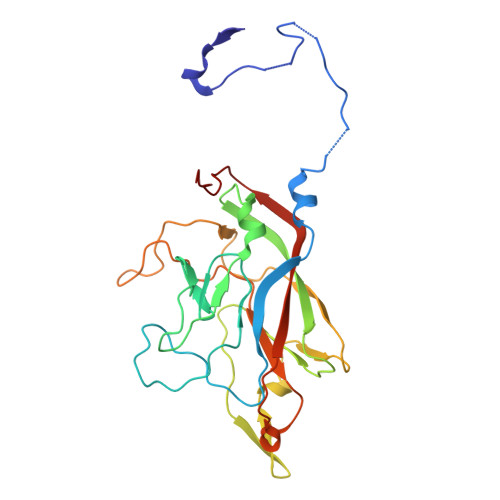
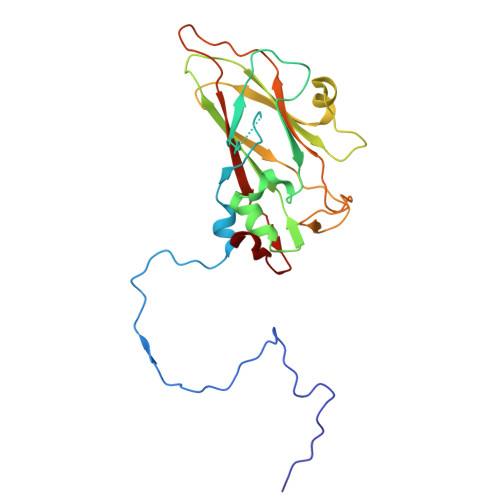
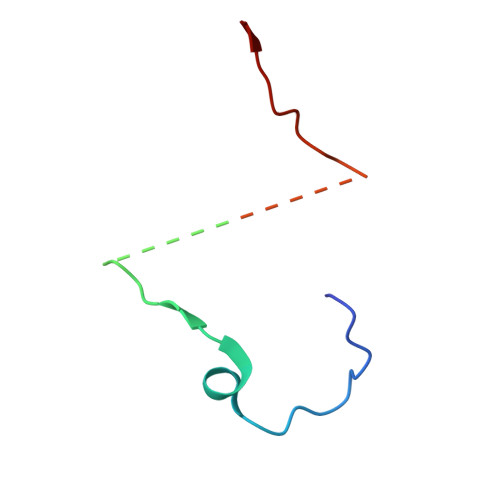
Cryo-EM structure of the Seneca Valley virus A-particle and related structural states.
Kumaran, R., Jayawardena, N., Chen, K.-.L., Eruera, A.-.R., Hodgkinson-Bean, J., Burga, L.N., Wolf, M., Bostina, M.(2025) J Virol 99: e0074425-e0074425
- PubMed: 40833065 Search on PubMed
- DOI: https://doi.org/10.1128/jvi.00744-25
- Primary Citation Related Structures:
9EAA, 9EAB, 9EAC, 9EAD - PubMed Abstract:
Picornavirus cell entry requires a series of capsid protein conformational changes leading to genome uncoating. For enteroviruses, receptor binding triggers the transition from a full (F) capsid to an altered (A) particle before releasing its genome and finally converting it into an empty (E) particle. In contrast, non-enteroviruses, such as Aphthovirus, Cardiovirus, or Seneca Valley virus, release their genomes by dissociating the capsid into pentamers. While the existence of a transient A-particle for non-enteroviruses was previously speculated, it has never been directly observed using structural methods. Seneca Valley virus (SVV) is an oncolytic picornavirus that selectively targets cancer cells by recognizing Tumor endothelial marker 8 (TEM8) as the host receptor. SVV disassembles into pentamers at acidic pH, suggesting that the acidic environment of the endosome could cause capsid disassembly. We used cryo-electron microscopy to investigate SVV under acidic conditions and in complex with TEM8 at physiological pH, identifying multiple uncoating intermediates. These include an altered-particle, an empty-rotated particle (E R ), and a series of open particles expelling the coiled genome. The A-particle is expanded, displays reduced interactions between capsid proteins, a reorganized genome, and has a poorly resolved VP1 N-terminus, VP2 N-terminus, and VP4. The E R particle has rotated pentamers, reduced contacts within the particle, lacks the genome, VP1 and VP2 N-termini, and VP4. Our work provides an understanding of transient SVV structural states and supports the existence of an intermediate SVV A-particle. These findings could help optimize SVV for oncolytic therapy.IMPORTANCESeneca Valley virus (SVV) is a non-enterovirus picornavirus with specific tumor tropism mediated by the receptor Tumor endothelial marker 8, also known as Anthrax toxin receptor 1. Using cryo-electron microscopy, it was possible to identify multiple structural states of SVV. We demonstrate that SVV capsids transition from full particles to altered (A) particles and then to empty-rotated (E R ) particles, with receptor binding and acidic pH driving these conformational changes, respectively. This study also identifies open particles with expelled genomes. Comparisons between A- and E R -particles reveal that peptide segments of VP1, VP2, and VP4 could potentially play a role in genome delivery. Future work can explore the formation of these structural states in vivo .
- Department of Microbiology and Immunology, University of Otago, Dunedin, Otago, New Zealand.
Organizational Affiliation: