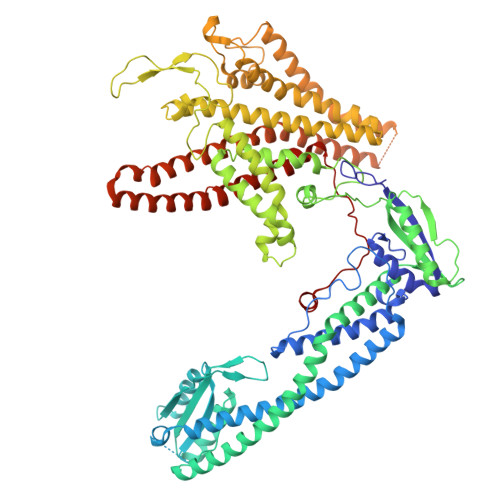
Monoclonal nanobodies alter the activity and assembly of the yeast vacuolar H + -ATPase.
Knight, K., Park, J.B., Oot, R.A., Khan, M.M., Roh, S.H., Wilkens, S.(2025) bioRxiv
- PubMed: 39829782 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.01.10.632502
- Primary Citation Related Structures:
9E76, 9E7L, 9MJ4 - PubMed Abstract:
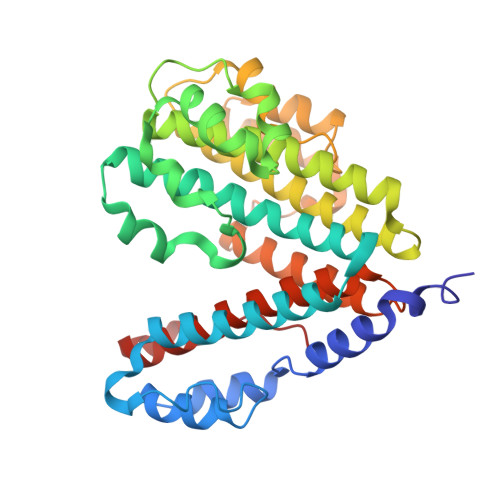
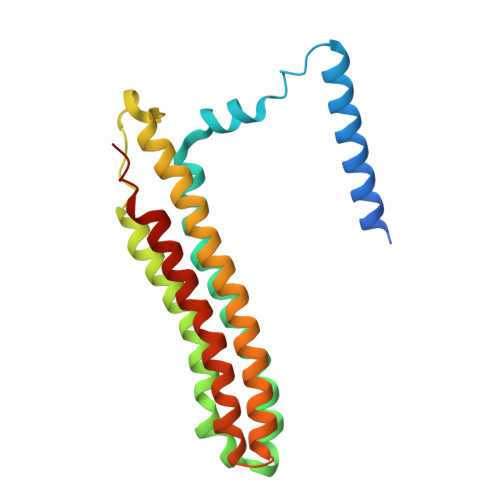
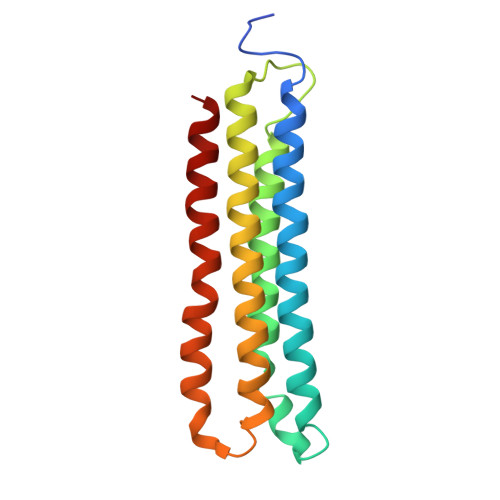
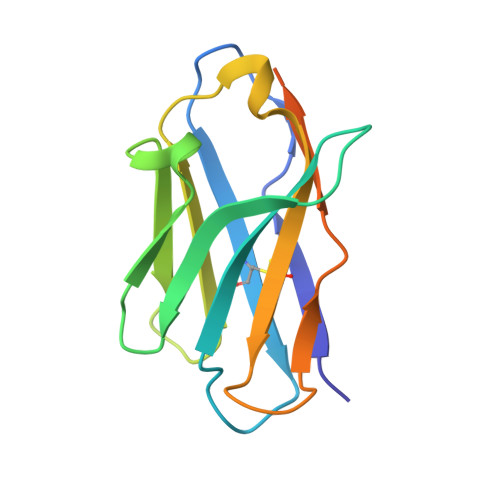
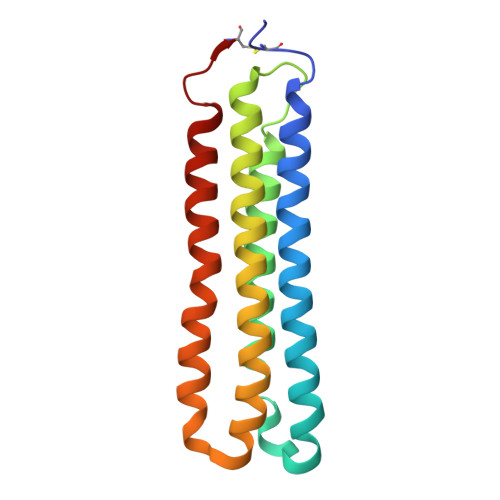
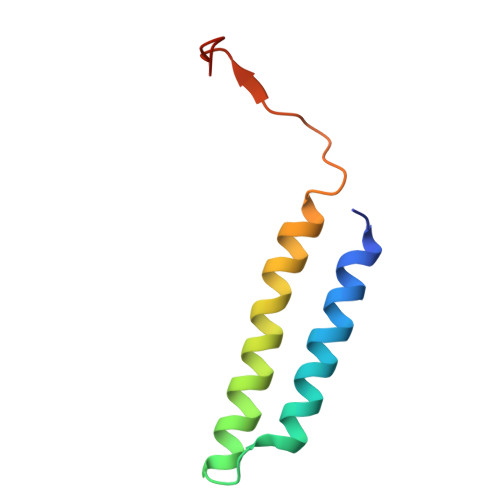
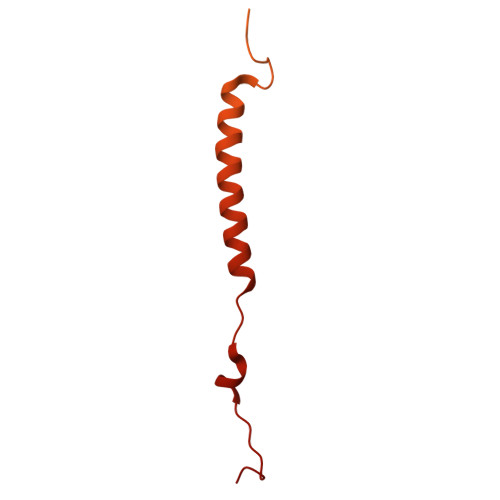
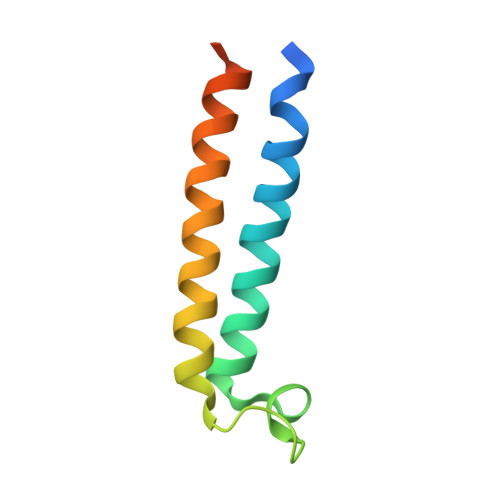
The vacuolar ATPase (V-ATPase; V 1 V o ) is a multi-subunit rotary nanomotor proton pump that acidifies organelles in virtually all eukaryotic cells, and extracellular spaces in some specialized tissues of higher organisms. Evidence suggests that metastatic breast cancers mislocalize V-ATPase to the plasma membrane to promote cell survival and facilitate metastasis, making the V-ATPase a potential drug target. We have generated a library of camelid single-domain antibodies (Nanobodies; Nbs) against lipid-nanodisc reconstituted yeast V-ATPase V o proton channel subcomplex. Here, we present an in-depth characterization of three anti-V o Nbs using biochemical and biophysical in vitro experiments. We find that the Nbs bind V o with high affinity, with one Nb inhibiting holoenzyme activity and another one preventing enzyme assembly. Using cryoEM, we find that two of the Nbs bind the c subunit ring of the V o on the lumen side of the complex. Additionally, we show that one of the Nbs raised against yeast V o can pull down human V-ATPase ( Hs V 1 V o ). Our research demonstrates Nb versatility to target and modulate the activity of the V-ATPase, and highlights the potential for future therapeutic Nb development.
- Department of Biochemistry and Molecular Biology, SUNY Upstate Medical University, Syracuse, NY 13210, USA.
Organizational Affiliation: