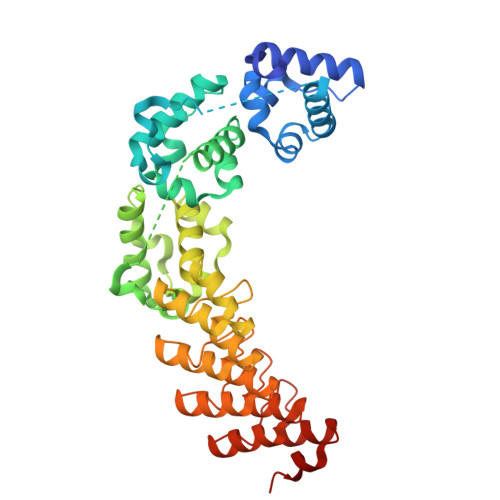
Optimal 1TEL-target protein linker character is target protein-dependent.
Pedroza Romo, M.J., Keliiliki, A., Averett, J.C., Gonzalez, J.F., Noakes, E., Wilson, E.W., Smith, C., Averett, B., Hansen, D., Nickles, R., Bradford, M., Soleimani, S., Smith, T., Nawarathnage, S., Samarwickrama, P., Kelsch, A., Bunn, D., Stewart, C., Abiodun, W., Tsubaki, E., Brown, S., Doukov, T.I., Moody, J.D.(2026) Acta Crystallogr D Struct Biol 82: 516-532
- PubMed: 41945409 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798326002494
- Primary Citation Related Structures:
9DB5, 9DP8, 9DVG, 9E4Q, 9ZNB - PubMed Abstract:
Fusing a variant of the sterile alpha motif domain of the human translocation ETS leukaemia protein (1TEL) to a protein of interest has been shown to significantly enhance its crystallization propensity. 1TEL is a pH-dependent, polymer-forming protein crystallization chaperone which, when covalently fused to a protein of interest, forms a stable, well ordered crystal lattice. However, despite its success, a challenge persists in that crystal quality and diffraction limits appear to be heavily dependent on the choice of linker between 1TEL and the protein of interest, with the identification of a functional linker currently relying on trial-and-error methods. Likewise, previous studies revealed that a ten-histidine tag at the 1TEL N-terminus can either facilitate or hinder the ordered crystallization of target proteins attached via flexible or semi-flexible linkers. To address these challenges, we designed multiple constructs with several types of linkers [rigid (helical fusion), semi-flexible (Pro-Ala and Pro-Ala-Ala) and flexible (Gly-Gly and Gly-Gly-Gly)] of varying lengths to fuse either a designed ankyrin-repeat protein (DARPin) or the thirty-eight-negative kinase-1 ubiquitin-associated (UBA) domain to the 1TEL C-terminus. Semi-flexible and flexible linker constructs were made with and without a ten-histidine tag. Our findings indicate that short semi-flexible and rigid linkers consistently yielded large crystals with a DARPin target protein, but that flexible linkers performed best with a UBA-domain target protein. Removing the ten-histidine tag uniformly enhanced crystallization rates, improved the crystal morphology and increased the crystallization propensity of the semi-flexible and flexible linker constructs. These results suggest that the ideal linker selection primarily depends on the properties of the target protein. Our data support our current recommendation to use a short flexible or semi-flexible linker between 1TEL and the target protein to facilitate protein crystallization and high-resolution structure determination.
- Department of Chemistry and Biochemistry, Brigham Young University, 701 East University Parkway, Provo, UT 84602, USA.
Organizational Affiliation: