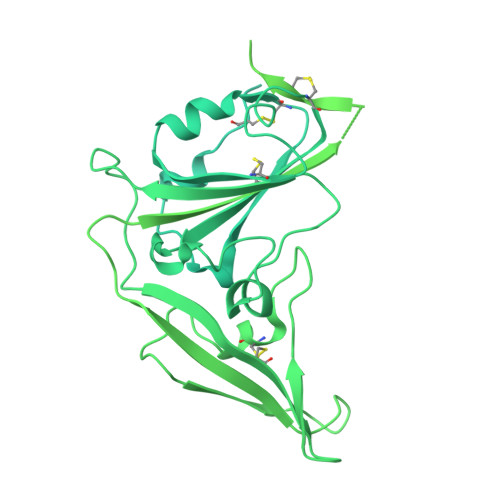
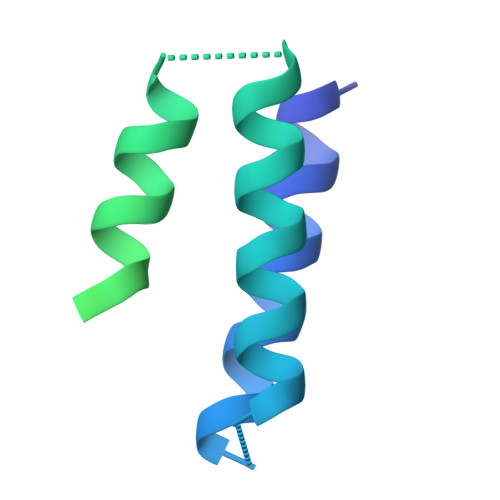
Designed miniproteins potently inhibit and protect against MERS-CoV.
Ragotte, R.J., Tortorici, M.A., Catanzaro, N.J., Addetia, A., Coventry, B., Froggatt, H.M., Lee, J., Stewart, C., Brown, J.T., Goreshnik, I., Sims, J.N., Milles, L.F., Wicky, B.I.M., Glogl, M., Gerben, S., Kang, A., Bera, A.K., Sharkey, W., Schafer, A., Harkema, J.R., Baric, R.S., Baker, D., Veesler, D.(2025) Cell Rep 44: 115760-115760
- PubMed: 40450691 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2025.115760
- Primary Citation Related Structures:
9C7Z, 9DGO, 9DKK - PubMed Abstract:
Middle East respiratory syndrome coronavirus (MERS-CoV) is a zoonotic pathogen with a 36% case-fatality rate in humans. No vaccines or specific therapeutics are currently approved for use in humans or the camel host reservoir. Here, we computationally designed monomeric and homo-oligomeric miniproteins that bind with high affinity to the MERS-CoV spike (S) glycoprotein, the main target of neutralizing antibodies and vaccine development. We show that these miniproteins broadly neutralize a panel of MERS-CoV S variants, spanning the known antigenic diversity of this pathogen, by targeting a conserved site in the receptor-binding domain (RBD). The miniproteins directly compete with binding of the dipeptidylpeptidase 4 (DPP4) receptor to MERS-CoV S, thereby blocking viral attachment to the host entry receptor and subsequent membrane fusion. Intranasal administration of a lead miniprotein provides prophylactic protection against stringent MERS-CoV challenge in mice, motivating its future clinical development as a next-generation countermeasure against this virus with pandemic potential.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA; Institute for Protein Design, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: