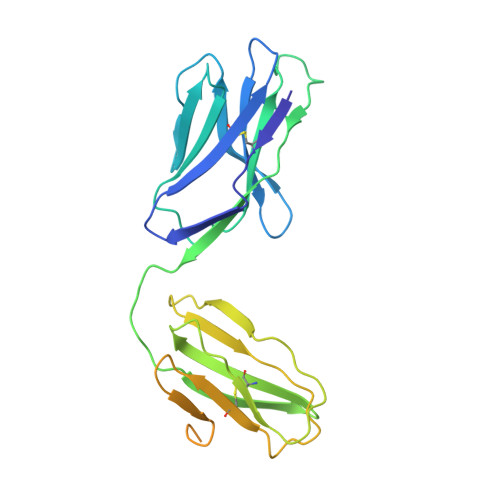
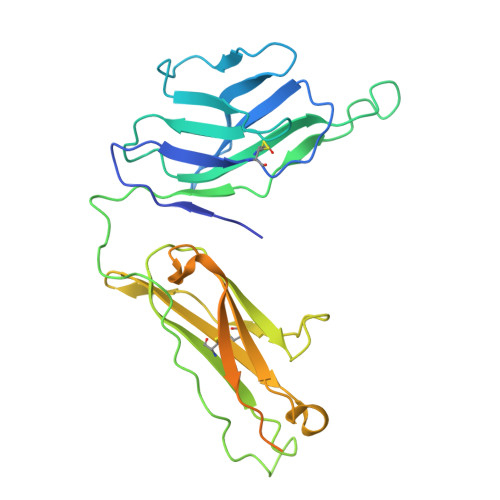
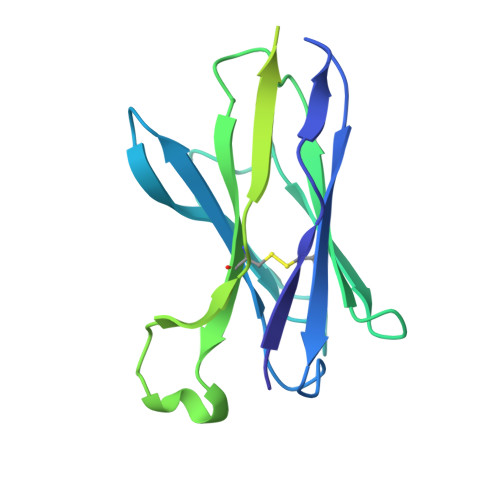
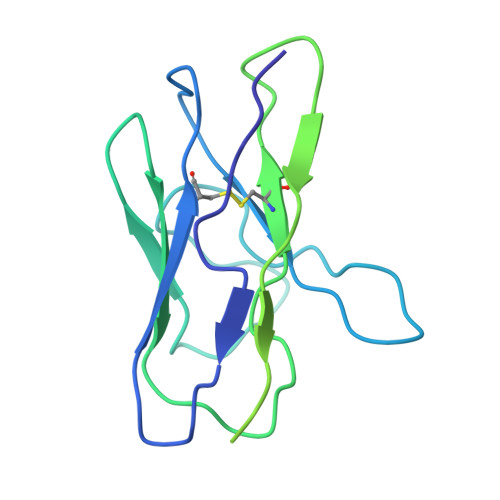
Structural characterization of two gamma delta TCR/CD3 complexes.
Hoque, M., Grigg, J.B., Ramlall, T., Jones, J., McGoldrick, L.L., Lin, J.C., Olson, W.C., Smith, E., Franklin, M.C., Zhang, T., Saotome, K.(2025) Nat Commun 16: 318-318
- PubMed: 39747888 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-55467-5
- Primary Citation Related Structures:
9CQ4, 9CQ7, 9CQ8, 9CQL - PubMed Abstract:
The T-cell receptor (TCR)/CD3 complex plays an essential role in the immune response and is a key player in cancer immunotherapies. There are two classes of TCR/CD3 complexes, defined by their TCR chain usage (αβ or γδ). Recently reported structures have revealed the organization of the αβ TCR/CD3 complex, but similar studies regarding the γδ TCR/CD3 complex have lagged behind. Here, we report cryoelectron microscopy (cryoEM) structural analysis of two γδ TCRs, G115 (Vγ9 Vδ2) and 9C2 (Vγ5 Vδ1), in complex with CD3 subunits. Our results show that the overall subunit organization of the γδ TCR/CD3 complexes is similar to αβ TCRs. However, both γδ TCRs display highly mobile extracellular domains (ECDs), unlike αβ TCRs, which have TCR ECDs that are rigidly coupled to its transmembrane (TM) domains. We corroborate this finding in cells by demonstrating that a γδ T-cell specific antibody can bind a site that would be inaccessible in the more rigid αβ TCR/CD3 complex. Furthermore, we observed that the Vγ5 Vδ1 complex forms a TCR γ5 chain-mediated dimeric species whereby two TCR/CD3 complexes are assembled. Collectively, these data shed light on γδ TCR/CD3 complex formation and may aid the design of γδ TCR-based therapies.
- Regeneron Pharmaceuticals, Inc., Tarrytown, NY, 10591, USA. mohammed.hoque@regeneron.com.
Organizational Affiliation: