Protective antibodies target cryptic epitope unmasked by cleavage of malaria sporozoite protein.
Dacon, C., Moskovitz, R., Swearingen, K., Da Silva Pereira, L., Flores-Garcia, Y., Aleshnick, M., Kanatani, S., Flynn, B., Molina-Cruz, A., Wollenberg, K., Traver, M., Kirtley, P., Purser, L., Dillon, M., Bonilla, B., Franco, A., Petros, S., Kritzberg, J., Tucker, C., Paez, G.G., Gupta, P., Shears, M.J., Pazzi, J., Edgar, J.M., Teng, A.A., Belmonte, A., Oda, K., Doumbo, S., Krymskaya, L., Skinner, J., Li, S., Ghosal, S., Kayentao, K., Ongoiba, A., Vaughan, A., Campo, J.J., Traore, B., Barillas-Mury, C., Wijayalath, W., Idris, A., Crompton, P.D., Sinnis, P., Wilder, B.K., Zavala, F., Seder, R.A., Wilson, I.A., Tan, J.(2025) Science 387: eadr0510-eadr0510
- PubMed: 39745947 Search on PubMed
- DOI: https://doi.org/10.1126/science.adr0510
- Primary Citation Related Structures:
9C79, 9C7D, 9C7F - PubMed Abstract:
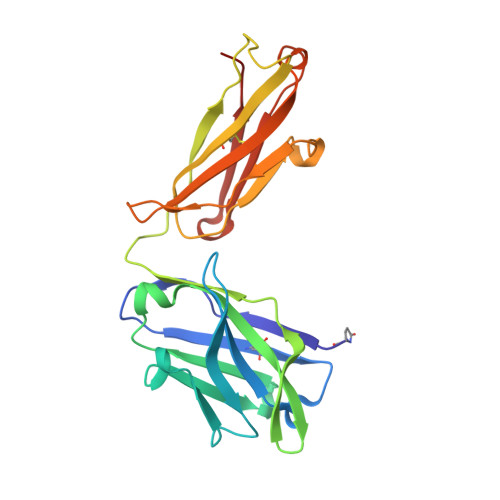
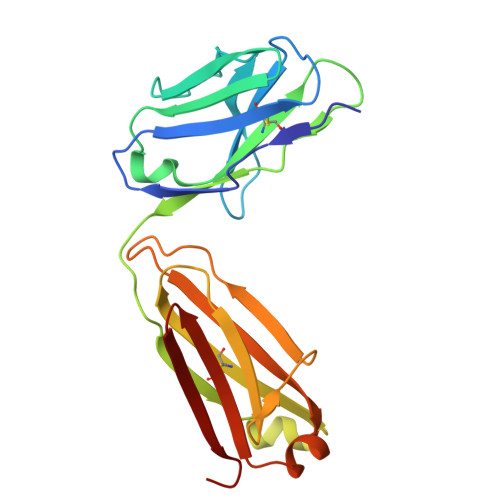
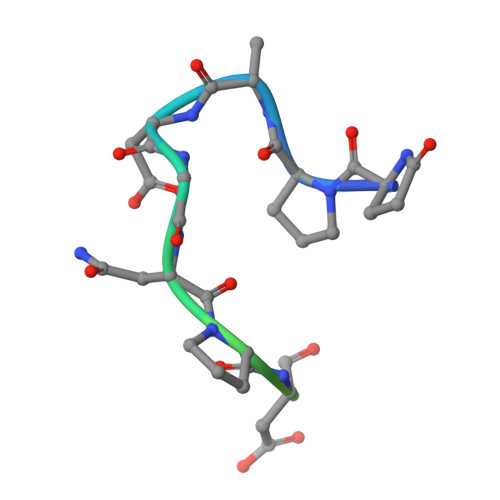
The most advanced monoclonal antibodies (mAbs) and vaccines against malaria target the central repeat region or closely related sequences within the Plasmodium falciparum circumsporozoite protein (PfCSP). Here, using an antigen-agnostic strategy to investigate human antibody responses to whole sporozoites, we identified a class of mAbs that target a cryptic PfCSP epitope that is only exposed after cleavage and subsequent pyroglutamylation (pGlu) of the newly formed N terminus. This pGlu-CSP epitope is not targeted by current anti-PfCSP mAbs and is not included in the licensed malaria vaccines. MAD21-101, the most potent mAb in this class, confers sterile protection against Pf infection in a human liver-chimeric mouse model. These findings reveal a site of vulnerability on the sporozoite surface that can be targeted by next-generation antimalarial interventions.
- Antibody Biology Unit, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Rockville, MD, USA.
Organizational Affiliation: