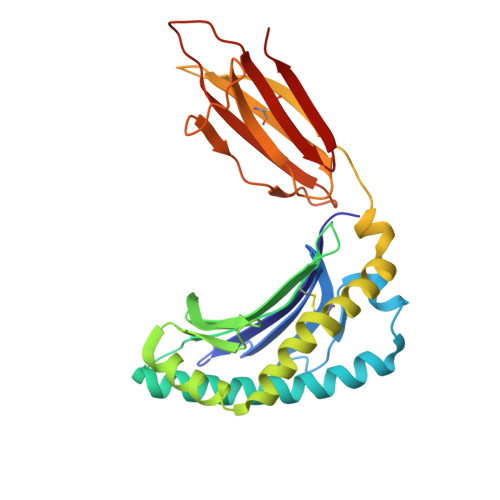
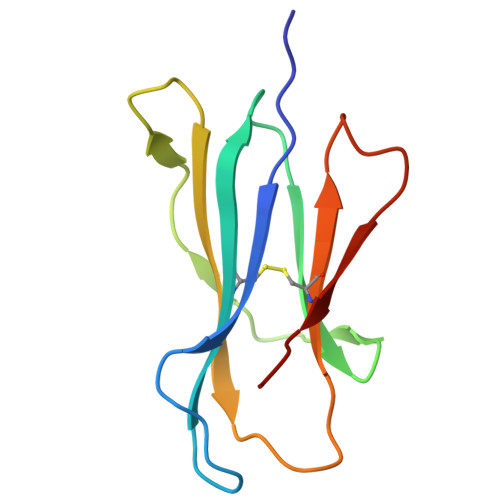
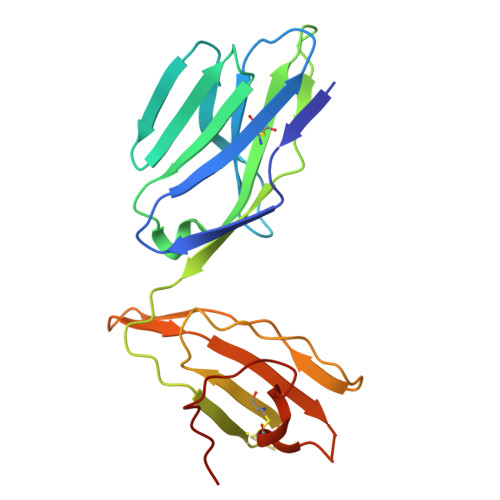
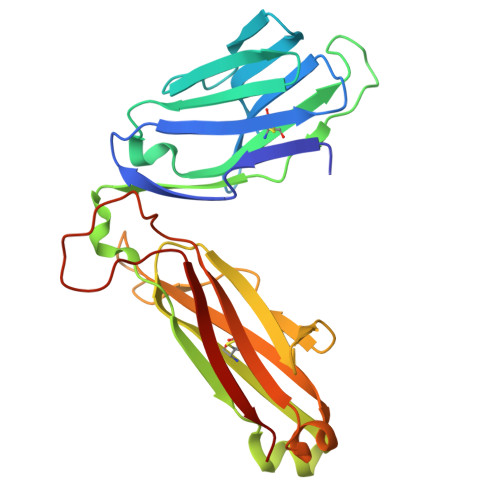
Ellagic acid metabolism as a source of dietary MR1 ligands.
O'Sullivan, D., Wang, C.J.H., Tang, J.S., Stephens, R., Faulkner, S., Gell, K., Li, Y., Cait, A., McInerney, M., Purcell, A.W., Rossjohn, J., Le Nours, J., Gasser, O.(2026) J Immunol 215
- PubMed: 41758608 Search on PubMed
- DOI: https://doi.org/10.1093/jimmun/vkaf346
- Primary Citation Related Structures:
9C42 - PubMed Abstract:
MR1 is an major histocompatibility complex class I-like molecule that presents small molecule metabolites to MR1-restricted T cells that include a major population of highly conserved T cells known as mucosal-associated invariant T (MAIT) cells. MAIT cells recognize bacterial riboflavin pathway-derived neoantigens and are being attributed an increasing number of immune and homeostatic functions. However, the chemical breadth and diversity of MR1-restricted ligands remain to be fully elucidated. Due to the largely (poly)cyclic structure of known MR1 ligands, we aimed to identify MR1 ligands from a library of dietary phenolic metabolites. Competitive MAIT cell inhibition assays using both cell lines and primary cells isolated from human blood identified gut microbial metabolites of ellagitannins that include ellagic acid (EA), urolithin D (UroD), and UroM5 as potential MR1 ligands. Fluorescence polarization binding assays demonstrated that EA, UroM5, UroC, and UroB bound to MR1, and we provide a structural basis for EA presentation by MR1. Overall, our findings indicate that EA metabolism provides dietary MR1 ligands that inhibit T cell receptor-dependent MAIT cell activation.
- Malaghan Institute of Medical Research, Wellington, New Zealand.
Organizational Affiliation: