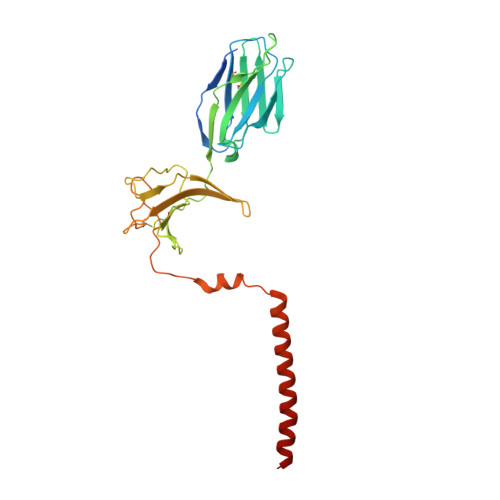
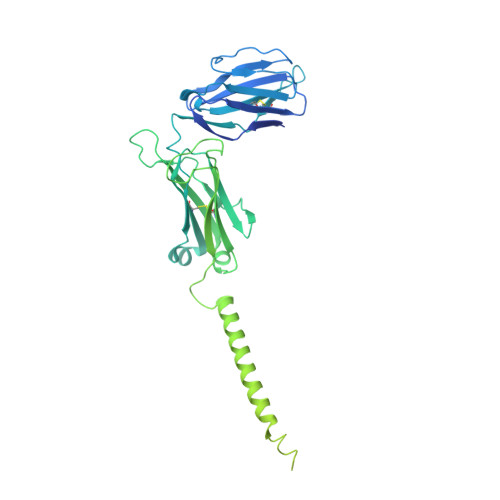
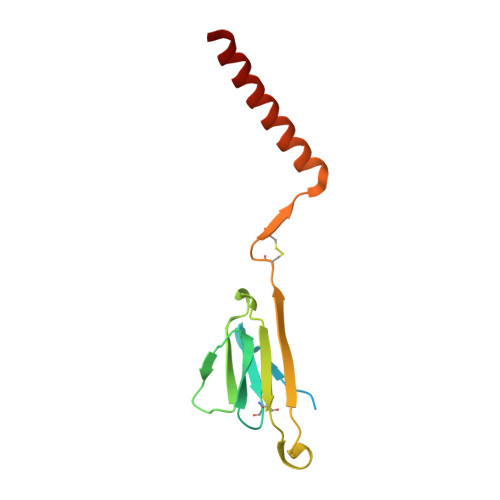
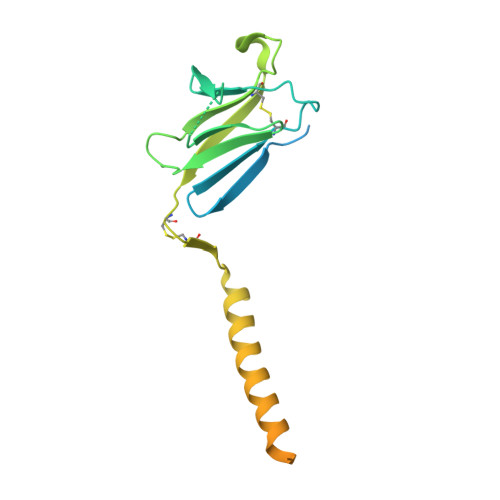
The resting and ligand-bound states of the membrane-embedded human T-cell receptor-CD3 complex.
Notti, R.Q., Yi, F., Heissel, S., Bush, M.W., Molvi, Z., Das, P., Molina, H., Klebanoff, C.A., Walz, T.(2025) Nat Commun 16: 10996-10996
- PubMed: 41402338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-66939-7
- Primary Citation Related Structures:
9BBC, 9C3E - PubMed Abstract:
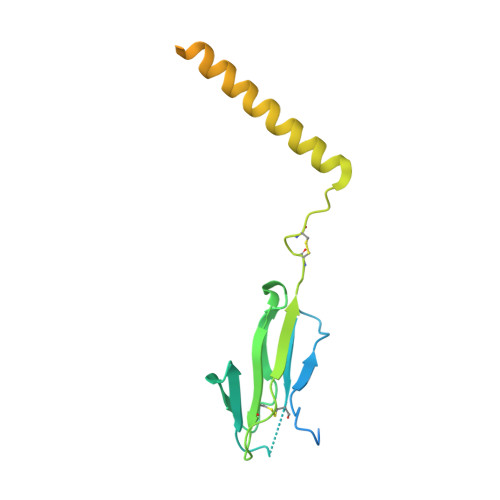
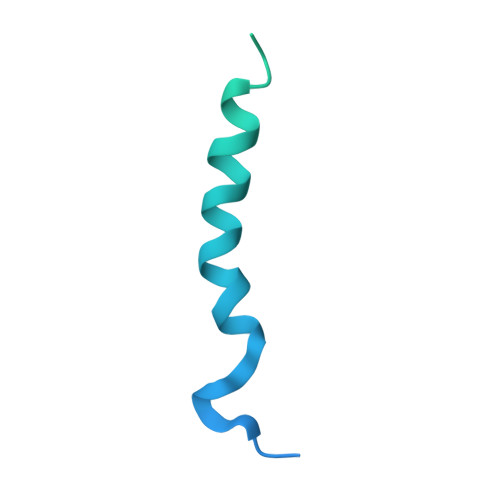
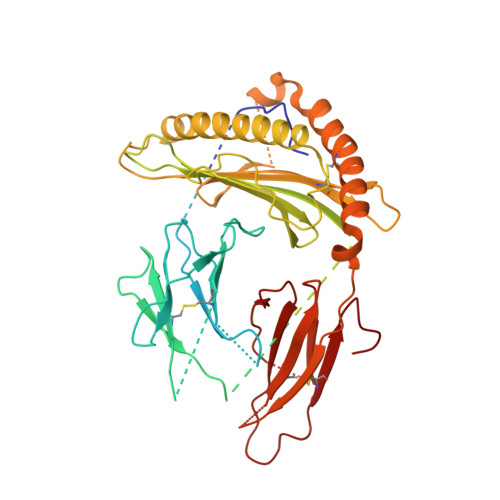
The T-cell receptor (TCR) initiates T-lymphocyte activation, but the mechanism of TCR activation remains uncertain. Here, we present cryogenic electron microscopy structures for the unliganded and human leukocyte antigen (HLA)-bound human TCR-CD3 complex in nanodiscs that provide a native-like lipid environment. Distinct from the open and extended conformation seen in detergent, the unliganded TCR-CD3 in nanodiscs adopts two related closed and compacted conformations that represent its physiologic resting state in vivo. By contrast, the HLA-bound complex adopts the open and extended conformation, and conformation-locking disulfide mutants show that ectodomain opening is necessary for maximal ligand-dependent T-cell activation. These structures also reveal conformation-dependent protein-lipid and glycan-glycan interactions within the TCR. Together, these results establish allosteric conformational change during TCR activation, reveal avenues for immunotherapeutic engineering, and highlight the importance of native-like lipid environments for membrane protein structure determination.
- Laboratory of Molecular Electron Microscopy, The Rockefeller University, New York, NY, USA. rnotti@rockefeller.edu.
Organizational Affiliation: