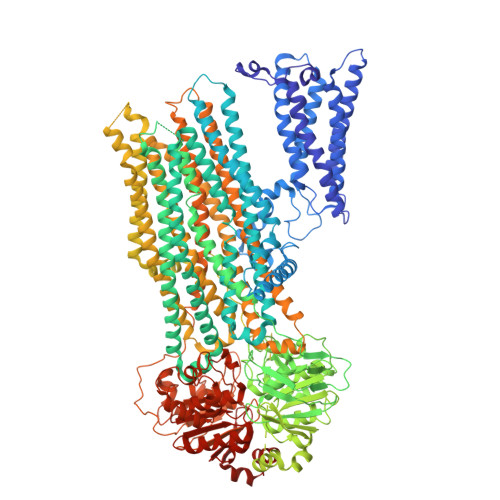
Structural basis for the transport and regulation mechanism of the multidrug resistance-associated protein 2.
Koide, E., Pietz, H.L., Beltran, J., Chen, J.(2025) Nat Commun 16: 484-484
- PubMed: 39779684 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-55810-w
- Primary Citation Related Structures:
9BR2, 9BUK, 9C12, 9C2I - PubMed Abstract:
Multidrug resistance-associated protein 2 (MRP2) is an ATP-powered exporter important for maintaining liver homeostasis and a potential contributor to chemotherapeutic resistance. Using cryogenic electron microscopy (cryo-EM), we determine the structures of human MRP2 in three conformational states: an autoinhibited state, a substrate-bound pre-translocation state, and an ATP-bound post-translocation state. In the autoinhibited state, the cytosolic regulatory (R) domain plugs into the transmembrane substrate-binding site and extends into the cytosol to form a composite ATP-binding site at the surface of nucleotide-binding domain 2. Substrate displaces the R domain, permitting conformational changes necessary for transport. These observations suggest that the R domain functions as a selectivity gauge, where only at sufficiently high concentrations can the substrate effectively initiate transport. Comparative structural analyzes of MRP2 bound to various substrates, as determined in this study and others, reveal how MRP2 recognizes a diverse array of compounds, supporting its role in multidrug resistance.
- Laboratory of Membrane Biology and Biophysics, The Rockefeller University, New York, NY, USA.
Organizational Affiliation: