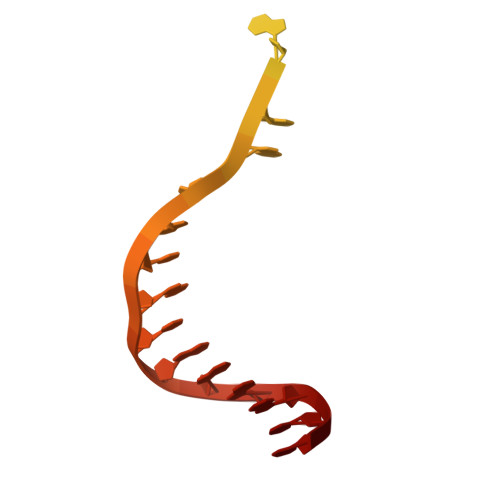
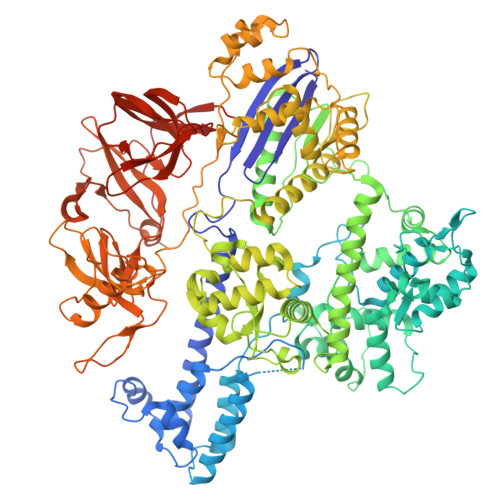
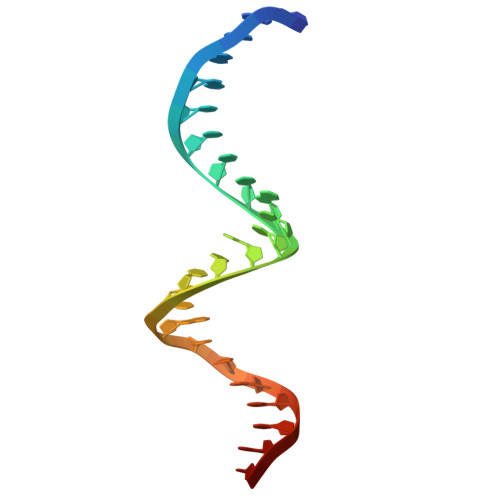
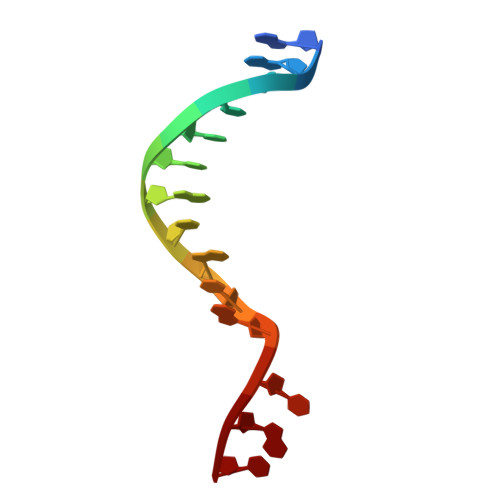
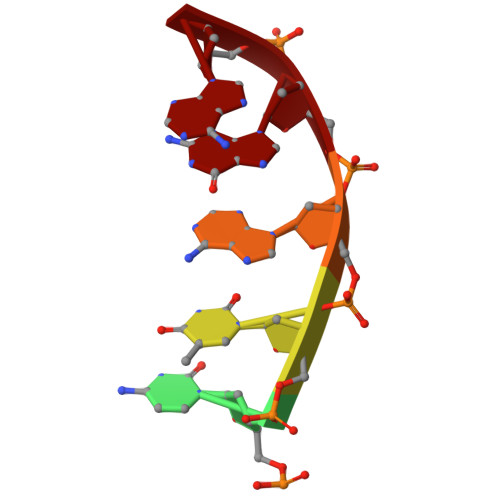
Molecular basis for methylation-sensitive editing by Cas9.
Roth, M.O., Shu, Y., Zhao, Y., Trasanidou, D., Hoffman, R.D., Sudfeld, C., Bouzetos, E., Trasanidis, N., Zawrotny, M., Gelasco, M.K., Medina, M.L., Das, A., Rai, J., Goswami, H.N., Wang, B., van der Oost, J., Li, H.(2026) Nature
- PubMed: 41986708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-026-10384-z
- Primary Citation Related Structures:
9BS6 - PubMed Abstract:
The bacterial CRISPR-Cas9 (Cas9) nuclease has become a powerful genome manipulation tool for a wide range of organisms 1-3 . However, it has yet to fully leverage the pervasive presence of DNA methylation in genomes 4-10 . Here, to fill this gap, we report biochemical, structural and human genome-editing characterizations of a methylation-sensitive Cas9 (ThermoCas9). ThermoCas9 efficiently binds to and cleaves DNA upstream of its protospacer adjacent motif (PAM) 5'-NNNNCGA-3' or 5'-NNNNCCA-3' in vitro. Methylation of the fifth cytosine in either PAM sequence ( 5m CpG or 5m CpC), however, significantly inhibits ThermoCas9 activity. Cryo-electron microscopy structures of ThermoCas9 in pre-cleavage and post-cleavage states at 2.8 Å and 2.2 Å resolution, respectively, reveal the molecular basis for the stringent requirement of the unmethylated cytosine in PAM binding and provide guidance for further enzyme engineering. We demonstrate methylation-sensitive editing by ThermoCas9 in human cell lines with distinct DNA methylation landscapes. Moreover, we demonstrate that a catalytically enhanced ThermoCas9 efficiently targets luminal expression signature genes that are consistently hypomethylated in patients with breast cancer. Owing to its sensitivity to DNA methylation, ThermoCas9 can specifically target cells with disease-related hypomethylation, which adds another layer of precision to genome-editing technologies.
- Department of Structural Biology, Van Andel Institute, Grand Rapids, MI, USA.
Organizational Affiliation: