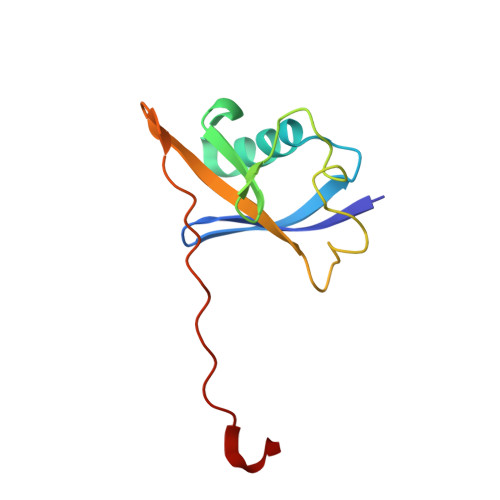
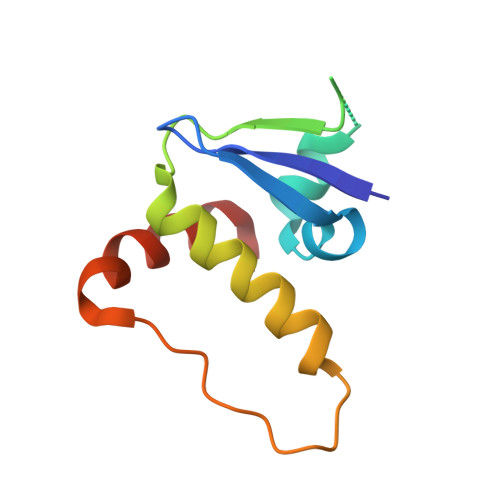
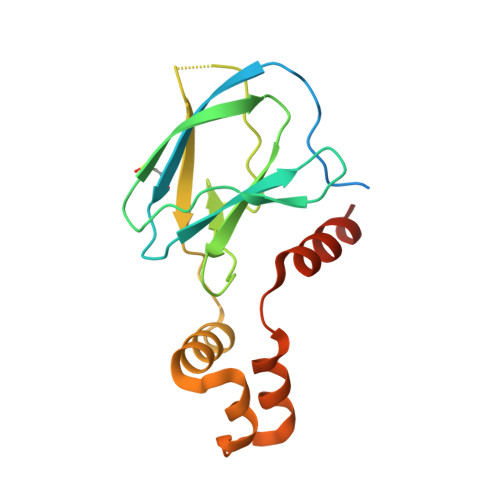
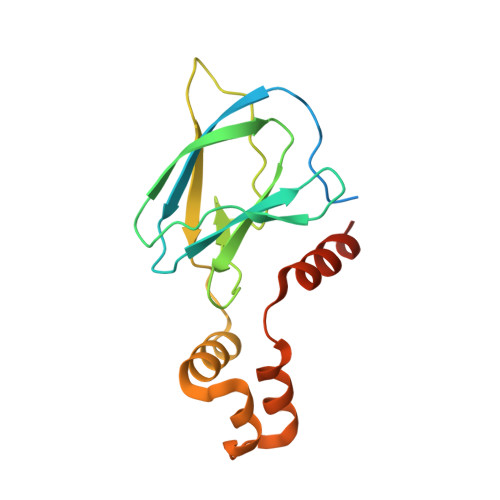
Potency-Enhanced Peptidomimetic VHL Ligands with Improved Oral Bioavailability.
Wu, H., Murray, J., Ishisoko, N., Frommlet, A., Deshmukh, G., DiPasquale, A., Mulvihill, M.M., Zhang, D., Quinn, J.G., Blake, R.A., Fairbrother, W.J., Fuhrmann, J.(2024) J Med Chem 67: 8585-8608
- PubMed: 38809766 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.3c02203
- Primary Citation Related Structures:
9BJU, 9BOL - PubMed Abstract:
The von Hippel-Lindau (VHL) protein plays a pivotal role in regulating the hypoxic stress response and has been extensively studied and utilized in the targeted protein degradation field, particularly in the context of bivalent degraders. In this study, we present a comprehensive peptidomimetic structure-activity relationship (SAR) approach, combined with cellular NanoBRET target engagement assays to enhance the existing VHL ligands. Through systematic modifications of the molecule, we identified the 1,2,3-triazole group as an optimal substitute of the left-hand side amide bond that yields 10-fold higher binding activity. Moreover, incorporating conformationally constrained alterations on the methylthiazole benzylamine moiety led to the development of highly potent VHL ligands with picomolar binding affinity and significantly improved oral bioavailability. We anticipate that our optimized VHL ligand, GNE7599 , will serve as a valuable tool compound for investigating the VHL pathway and advancing the field of targeted protein degradation.
- Department of Early Discovery Biochemistry, Genentech, 1 DNA Way, South San Francisco, California 94080, United States.
Organizational Affiliation: