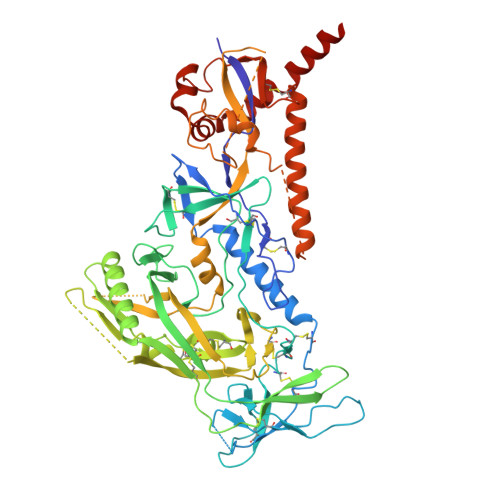
Accelerated cGMP production of near-native HIV-1 Env trimers following electroporation transfection and immunogenicity analysis.
Bale, S., Gustchina, E., Guenaga, J., Ayala, V., Lee, W.H., Ozorowski, G., Whitney, S., Wilson, R., Baboo, S., Diedrich, J.K., Doyle, E.D., Hudacik, L., Ben-Akiva, E., Rodrigues, K.A., Irvine, D.J., Yates 3rd, J.R., Paulson, J.C., Ward, A.B., Fouts, T., Wyatt, R.T.(2025) NPJ Vaccines 10: 198-198
- PubMed: 40835829 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41541-025-01218-6
- Primary Citation Related Structures:
9BE9 - PubMed Abstract:
Evaluation of recombinant HIV-1 surface glycoproteins (Env) as vaccine candidates for Phase I human experimental trials often requires production of cGMP-grade well-ordered Env trimers. Here, we report an accelerated cGMP compatible approach for expression and purification of a stabilized HIV clade C-derived trimer '16055 DG4 NFL' (for native flexibly linked). This recombinant trimer was expressed from CHO-S™ cells using a MaxCyte® VLX™ electroporation-based transient transfection process. The 16055 DG4 NFL was designed with multiple internal stabilizing mutations and, as well, deletion of four N-linked glycans (DG4) proximal to the CD4 binding site (CD4bs) engineered to improve B cell recognition of this conserved neutralizing determinant. The transient process circumvents the need to develop stable cell lines expressing the Env trimers that is often the most time-consuming step impacting vaccine development timelines. The 16055 DG4 NFL trimer was purified by immunoaffinity chromatography using the broadly neutralizing antibody (bNAb), PGT145. Following additional downstream processing steps, purified trimer was vialed, frozen and stored at -80 °C. Upon thaw and analysis, the trimer displayed homogeneity and a near-native conformation as determined by size-exclusion chromatography (SEC), negative stain and cryo-electron microscopy (EM), differential scanning calorimetry (DSC) and biolayer interferometry (BLI). The immunogenicity of the trimer was tested in rabbits with bolus, escalating dose and divided dose immunization regimens. Rabbits from all three regimens elicited tier 2 autologous neutralizing antibodies that targeted the exposed protein region at the CD4bs. The trimer is currently under investigation in a human clinical trial (NCT06332339) for safety, tolerability and as a priming candidate followed by heterologous boosting to potentially elicit cross-neutralizing antibodies.
- Department of Immunology and Microbiology, The Scripps Research Institute, La Jolla, CA, USA.
Organizational Affiliation: