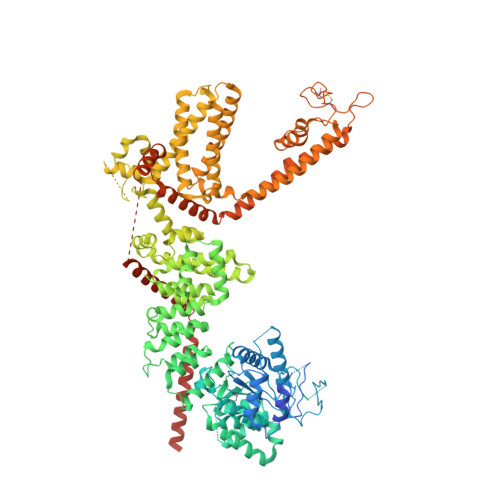
Mechanisms of sensory adaptation and inhibition of the cold and menthol receptor TRPM8.
Yin, Y., Park, C.G., Zhang, F., G Fedor, J., Feng, S., Suo, Y., Im, W., Lee, S.Y.(2024) Sci Adv 10: eadp2211-eadp2211
- PubMed: 39093967 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adp2211
- Primary Citation Related Structures:
9B6D, 9B6E, 9B6F, 9B6G, 9B6H, 9B6I, 9B6J, 9B6K - PubMed Abstract:
Our sensory adaptation to cold and chemically induced coolness is mediated by the intrinsic property of TRPM8 channels to desensitize. TRPM8 is also implicated in cold-evoked pain disorders and migraine, highlighting its inhibitors as an avenue for pain relief. Despite the importance, the mechanisms of TRPM8 desensitization and inhibition remained unclear. We found, using cryo-electron microscopy, electrophysiology, and molecular dynamics simulations, that TRPM8 inhibitors bind selectively to the desensitized state of the channel. These inhibitors were used to reveal the overlapping mechanisms of desensitization and inhibition and that cold and cooling agonists share a common desensitization pathway. Furthermore, we identified the structural determinants crucial for the conformational change in TRPM8 desensitization. Our study illustrates how receptor-level conformational changes alter cold sensation, providing insights into therapeutic development.
- Department of Biochemistry, Duke University School of Medicine, Durham, NC 27710, USA.
Organizational Affiliation: