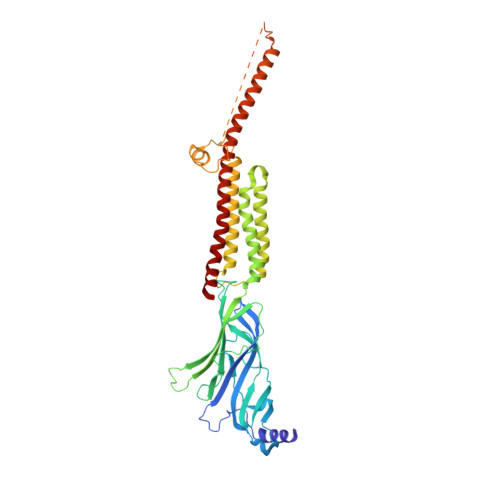
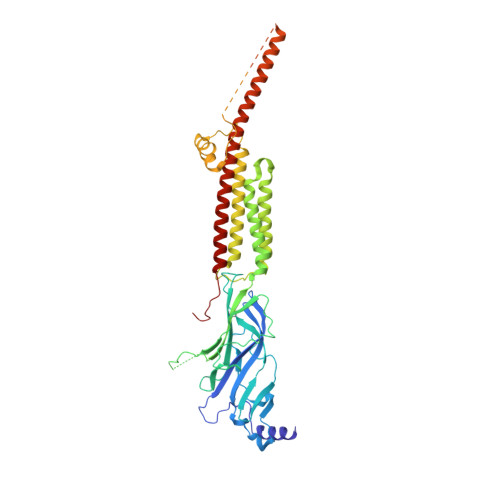
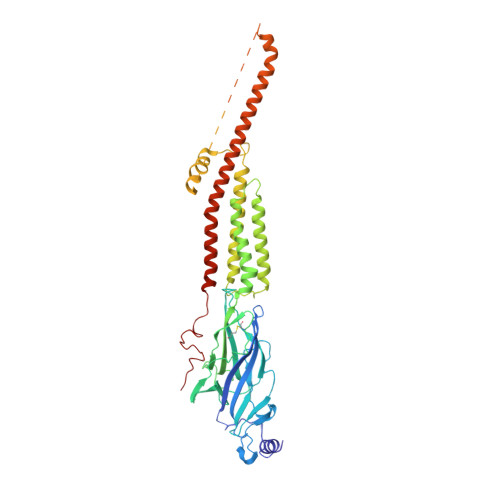
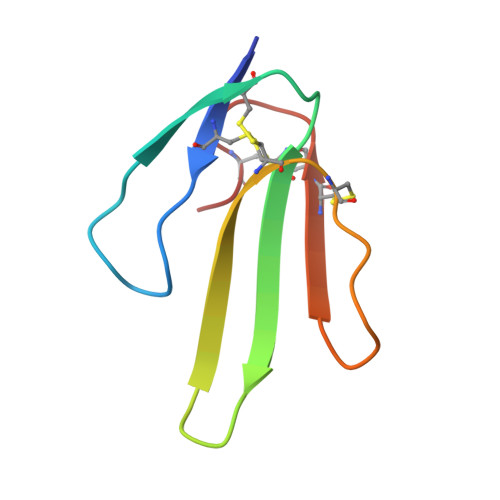
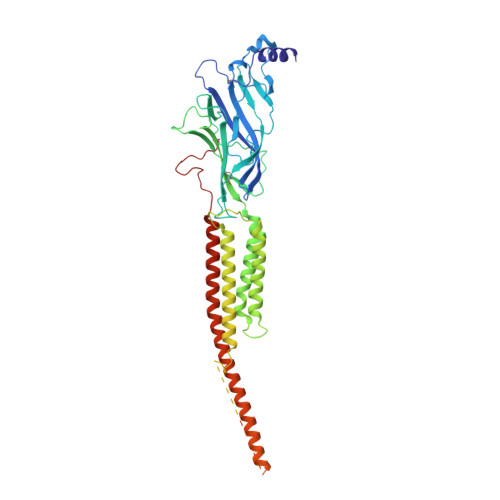
Structural switch in acetylcholine receptors in developing muscle.
Li, H., Teng, J., Hibbs, R.E.(2024) Nature 632: 1174-1180
- PubMed: 39085615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07774-6
- Primary Citation Related Structures:
9AVU, 9AVV, 9AWJ, 9AWK - PubMed Abstract:
During development, motor neurons originating in the brainstem and spinal cord form elaborate synapses with skeletal muscle fibres 1 . These neurons release acetylcholine (ACh), which binds to nicotinic ACh receptors (AChRs) on the muscle, initiating contraction. Two types of AChR are present in developing muscle cells, and their differential expression serves as a hallmark of neuromuscular synapse maturation 2-4 . The structural principles underlying the switch from fetal to adult muscle receptors are unknown. Here, we present high-resolution structures of both fetal and adult muscle nicotinic AChRs, isolated from bovine skeletal muscle in developmental transition. These structures, obtained in the absence and presence of ACh, provide a structural context for understanding how fetal versus adult receptor isoforms are tuned for synapse development versus the all-or-none signalling required for high-fidelity skeletal muscle contraction. We find that ACh affinity differences are driven by binding site access, channel conductance is tuned by widespread surface electrostatics and open duration changes result from intrasubunit interactions and structural flexibility. The structures further reveal pathogenic mechanisms underlying congenital myasthenic syndromes.
- Department of Neurobiology, University of California San Diego, La Jolla, CA, USA.
Organizational Affiliation: