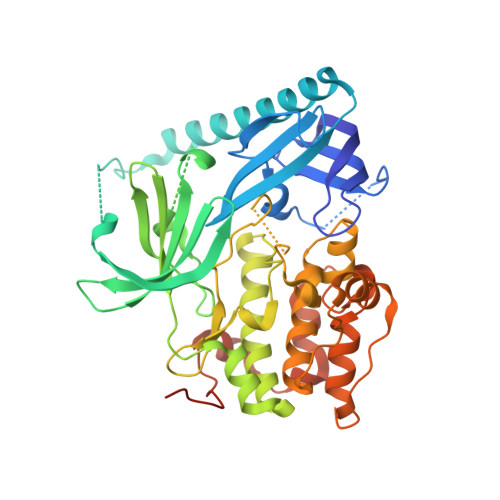
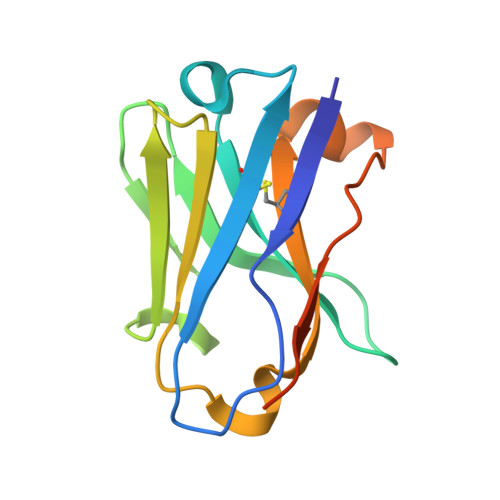
Mutant-selective AKT inhibition through lysine targeting and neo-zinc chelation.
Craven, G.B., Chu, H., Sun, J.D., Carelli, J.D., Coyne, B., Chen, H., Chen, Y., Ma, X., Das, S., Kong, W., Zajdlik, A.D., Yang, K.S., Reisberg, S.H., Thompson, P.A., Lipford, J.R., Taunton, J.(2025) Nature 637: 205-214
- PubMed: 39506119 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-08176-4
- Primary Citation Related Structures:
8UVY, 8UW2, 8UW7, 8UW9, 9C1W - PubMed Abstract:
Somatic alterations in the oncogenic kinase AKT1 have been identified in a broad spectrum of solid tumours. The most common AKT1 alteration replaces Glu17 with Lys (E17K) in the regulatory pleckstrin homology domain 1 , resulting in constitutive membrane localization and activation of oncogenic signalling. In clinical studies, pan-AKT inhibitors have been found to cause dose-limiting hyperglycaemia 2-6 , which has motivated the search for mutant-selective inhibitors. We exploited the E17K mutation to design allosteric, lysine-targeted salicylaldehyde inhibitors with selectivity for AKT1 (E17K) over wild-type AKT paralogues, a major challenge given the presence of three conserved lysines near the allosteric site. Crystallographic analysis of the covalent inhibitor complex unexpectedly revealed an adventitious tetrahedral zinc ion that coordinates two proximal cysteines in the kinase activation loop while simultaneously engaging the E17K-imine conjugate. The salicylaldimine complex with AKT1 (E17K), but not that with wild-type AKT1, recruits endogenous Zn 2+ in cells, resulting in sustained inhibition. A salicylaldehyde-based inhibitor was efficacious in AKT1 (E17K) tumour xenograft models at doses that did not induce hyperglycaemia. Our study demonstrates the potential to achieve exquisite residence-time-based selectivity for AKT1 (E17K) by targeting the mutant lysine together with Zn 2+ chelation by the resulting salicylaldimine adduct.
- Department of Cellular and Molecular Pharmacology, University of California, San Francisco, San Francisco, CA, USA.
Organizational Affiliation: