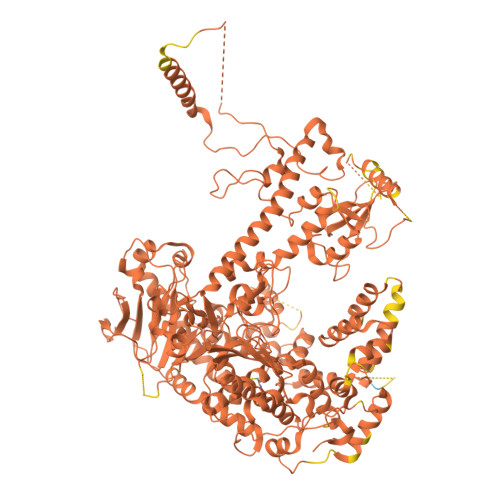
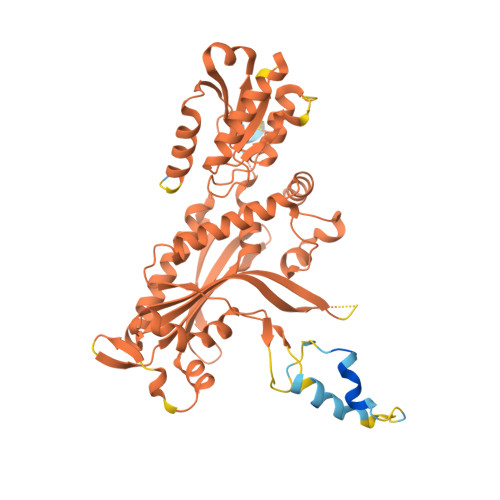
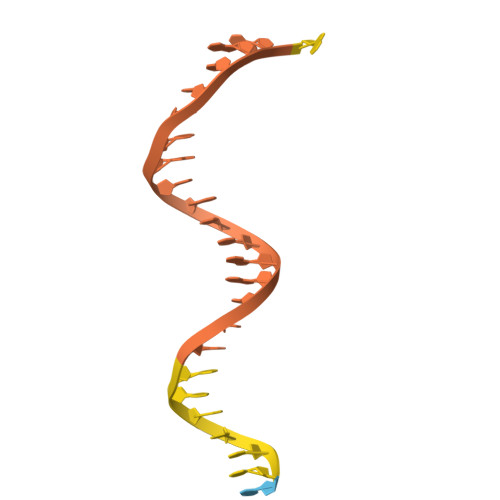
An interaction network in the polymerase active site is a prerequisite for Watson-Crick base pairing in Pol gamma.
Park, J., Herrmann, G.K., Roy, A., Shumate, C.K., Cisneros, G.A., Yin, Y.W.(2024) Sci Adv 10: eadl3214-eadl3214
- PubMed: 38787958 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adl3214
- Primary Citation Related Structures:
8UDK, 8UDL - PubMed Abstract:
The replication accuracy of DNA polymerase gamma (Pol γ) is essential for mitochondrial genome integrity. Mutation of human Pol γ arginine-853 has been linked to neurological diseases. Although not a catalytic residue, Pol γ arginine-853 mutants are void of polymerase activity. To identify the structural basis for the disease, we determined a crystal structure of the Pol γ mutant ternary complex with correct incoming nucleotide 2'-deoxycytidine 5'-triphosphate (dCTP). Opposite to the wild type that undergoes open-to-closed conformational changes when bound to a correct nucleotide that is essential for forming a catalytically competent active site, the mutant complex failed to undergo the conformational change, and the dCTP did not base pair with its Watson-Crick complementary templating residue. Our studies revealed that arginine-853 coordinates an interaction network that aligns the 3'-end of primer and dCTP with the catalytic residues. Disruption of the network precludes the formation of Watson-Crick base pairing and closing of the active site, resulting in an inactive polymerase.
- Department of Biochemistry and Molecular Biology, University of Texas Medical Branch, Galveston, TX 77555, USA.
Organizational Affiliation: