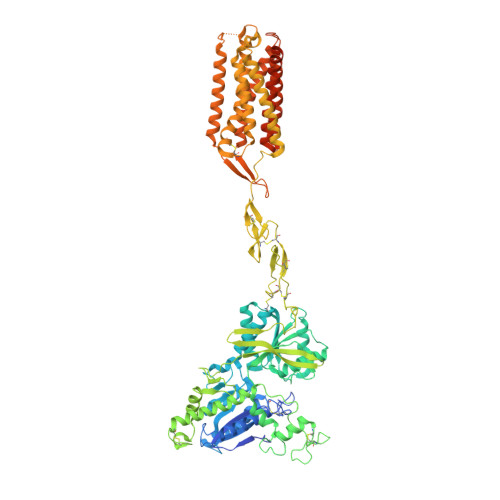
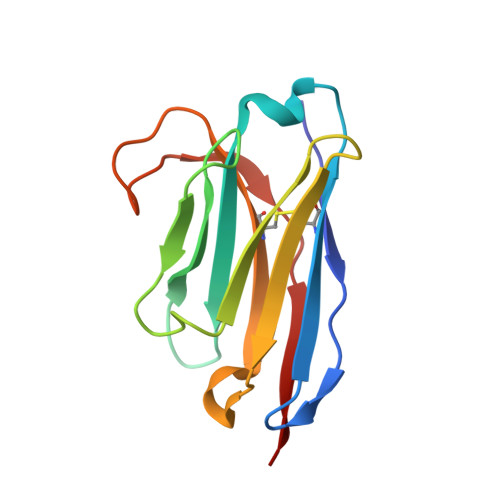
Stepwise activation of a metabotropic glutamate receptor.
Krishna Kumar, K., Wang, H., Habrian, C., Latorraca, N.R., Xu, J., O'Brien, E.S., Zhang, C., Montabana, E., Koehl, A., Marqusee, S., Isacoff, E.Y., Kobilka, B.K.(2024) Nature 629: 951-956
- PubMed: 38632403 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07327-x
- Primary Citation Related Structures:
8T6J, 8T7H, 8T8M, 8TAO - PubMed Abstract:
Metabotropic glutamate receptors belong to a family of G protein-coupled receptors that are obligate dimers and possess a large extracellular ligand-binding domain that is linked via a cysteine-rich domain to their 7-transmembrane domain 1 . Upon activation, these receptors undergo a large conformational change to transmit the ligand binding signal from the extracellular ligand-binding domain to the G protein-coupling 7-transmembrane domain 2 . In this manuscript, we propose a model for a sequential, multistep activation mechanism of metabotropic glutamate receptor subtype 5. We present a series of structures in lipid nanodiscs, from inactive to fully active, including agonist-bound intermediate states. Further, using bulk and single-molecule fluorescence imaging, we reveal distinct receptor conformations upon allosteric modulator and G protein binding.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA, USA. kaavyak@stanford.edu.
Organizational Affiliation: