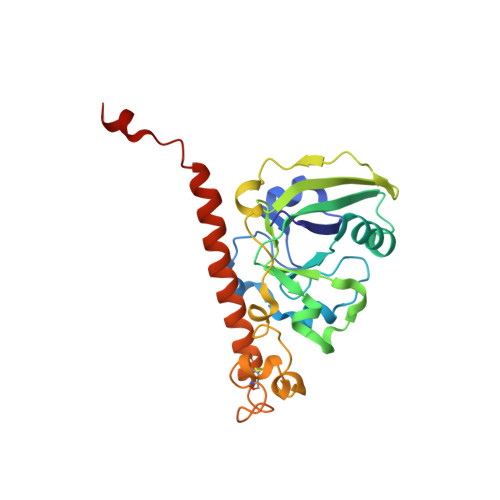
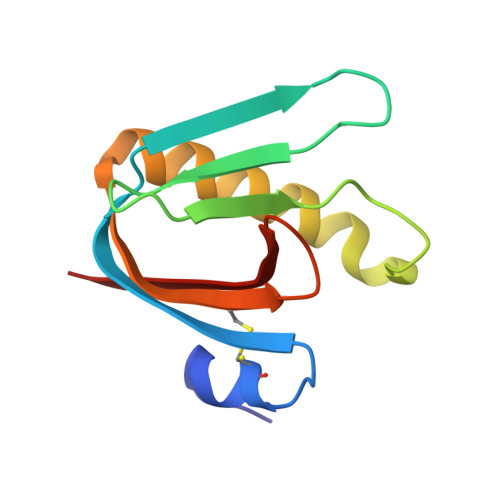
Using Vibrio natriegens for High-Yield Production of Challenging Expression Targets and for Protein Perdeuteration.
Mojica, N., Kersten, F., Montserrat-Canals, M., Huhn Iii, G.R., Tislevoll, A.M., Cordara, G., Teter, K., Krengel, U.(2024) Biochemistry 63: 587-598
- PubMed: 38359344 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.3c00612
- Primary Citation Related Structures:
8QRE - PubMed Abstract:
Production of soluble proteins is essential for structure/function studies; however, this usually requires milligram amounts of protein, which can be difficult to obtain with traditional expression systems. Recently, the Gram-negative bacterium Vibrio natriegens emerged as a novel and alternative host platform for production of proteins in high yields. Here, we used a commercial strain derived from V. natriegens (Vmax X2) to produce soluble bacterial and fungal proteins in milligram scale, which we struggled to achieve in Escherichia coli . These proteins include the cholera toxin (CT) and N -acetyl glucosamine-binding protein A (GbpA) from Vibrio cholerae , the heat-labile enterotoxin (LT) from E. coli and the fungal nematotoxin CCTX2 from Coprinopsis cinerea . CT, GbpA, and LT are secreted by the Type II secretion system in their natural hosts. When these three proteins were produced in Vmax, they were also secreted and could be recovered from the growth media. This simplified the downstream purification procedure and resulted in considerably higher protein yields compared to production in E. coli (6- to 26-fold increase). We also tested Vmax for protein perdeuteration using deuterated minimal media with deuterium oxide as solvent and achieved a 3-fold increase in yield compared to the equivalent protocol in E. coli . This is good news, since isotopic labeling is expensive and often ineffective but represents a necessary prerequisite for some structural biology techniques. Thus, Vmax represents a promising host for production of challenging expression targets and for protein perdeuteration in amounts suitable for structural biology studies.
- Department of Chemistry, University of Oslo, NO-0315 Blindern, Oslo, Norway.
Organizational Affiliation: