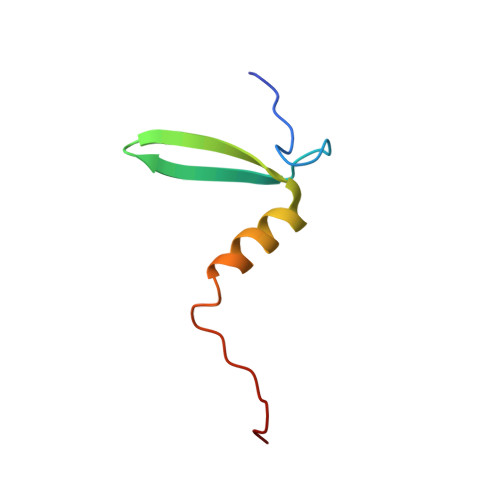
Structural basis of aggregate binding by the AAA+ disaggregase ClpG.
Katikaridis, P., Simon, B., Jenne, T., Moon, S., Lee, C., Hennig, J., Mogk, A.(2023) J Biological Chem 299: 105336-105336
- PubMed: 37827289 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.105336
- Primary Citation Related Structures:
8P66 - PubMed Abstract:
Severe heat stress causes massive loss of essential proteins by aggregation, necessitating a cellular activity that rescues aggregated proteins. This activity is executed by ATP-dependent, ring-forming, hexameric AAA+ disaggregases. Little is known about the recognition principles of stress-induced protein aggregates. How can disaggregases specifically target aggregated proteins, while avoiding binding to soluble non-native proteins? Here, we determined by NMR spectroscopy the core structure of the aggregate-targeting N1 domain of the bacterial AAA+ disaggregase ClpG, which confers extreme heat resistance to bacteria. N1 harbors a Zn 2+ -coordination site that is crucial for structural integrity and disaggregase functionality. We found that conserved hydrophobic N1 residues located on a β-strand are crucial for aggregate targeting and disaggregation activity. Analysis of mixed hexamers consisting of full-length and N1-truncated subunits revealed that a minimal number of four N1 domains must be present in a AAA+ ring for high-disaggregation activity. We suggest that multiple N1 domains increase substrate affinity through avidity effects. These findings define the recognition principle of a protein aggregate by a disaggregase, involving simultaneous contacts with multiple hydrophobic substrate patches located in close vicinity on an aggregate surface. This binding mode ensures selectivity for aggregated proteins while sparing soluble, non-native protein structures from disaggregase activity.
- Center for Molecular Biology of Heidelberg University (ZMBH), DKFZ-ZMBH Alliance, Heidelberg, Germany; German Cancer Research Center (DKFZ), Heidelberg, Germany.
Organizational Affiliation: