Detection of a Geminate Photoproduct of Bovine Cytochrome c Oxidase by Time-Resolved Serial Femtosecond Crystallography.
Ishigami, I., Carbajo, S., Zatsepin, N., Hikita, M., Conrad, C.E., Nelson, G., Coe, J., Basu, S., Grant, T., Seaberg, M.H., Sierra, R.G., Hunter, M.S., Fromme, P., Fromme, R., Rousseau, D.L., Yeh, S.R.(2023) J Am Chem Soc 145: 22305-22309
- PubMed: 37695261 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.3c07803
- Primary Citation Related Structures:
8GBT - PubMed Abstract:
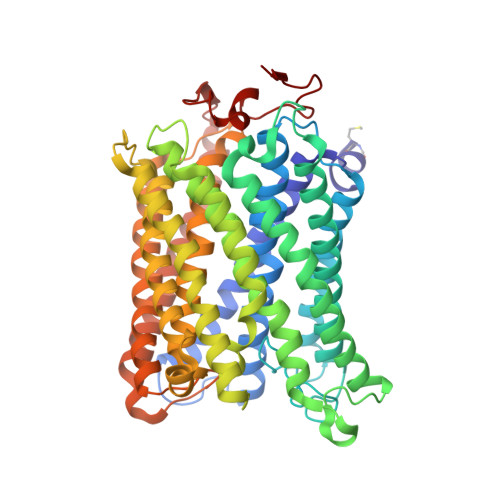
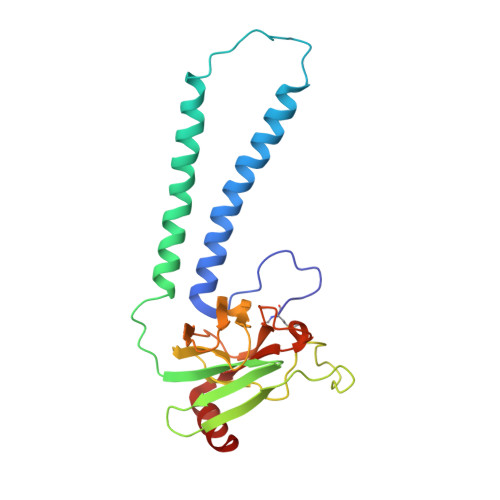
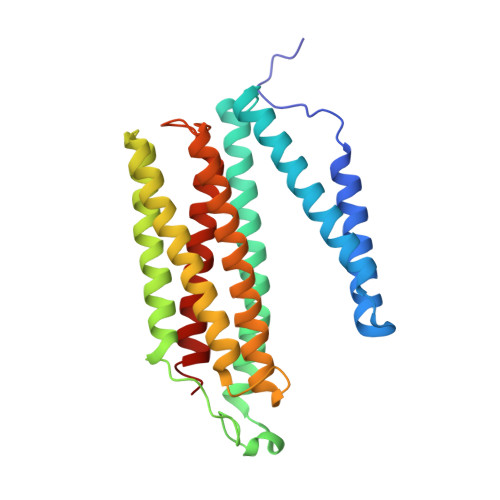
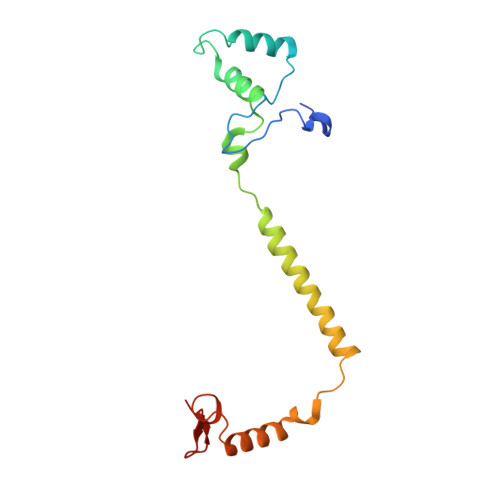
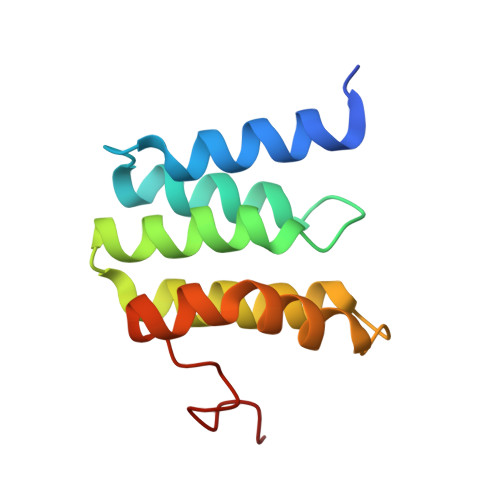
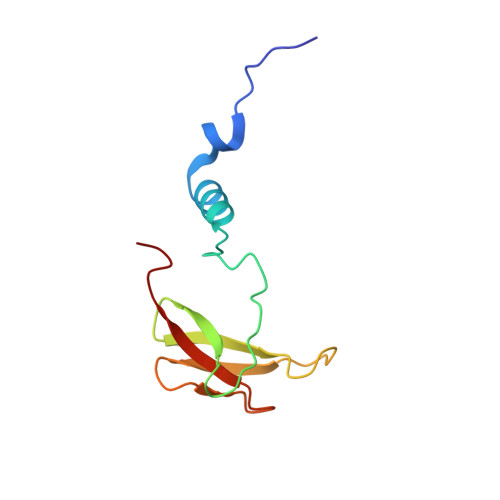
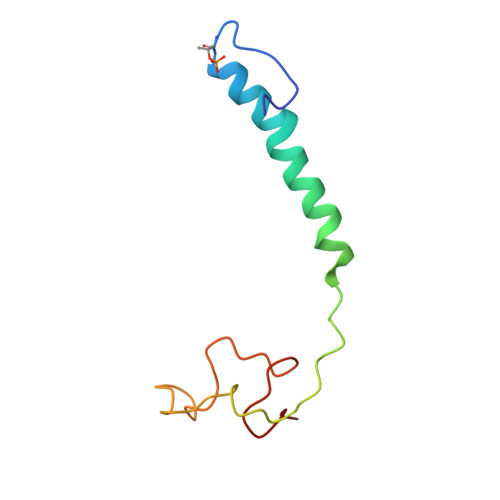
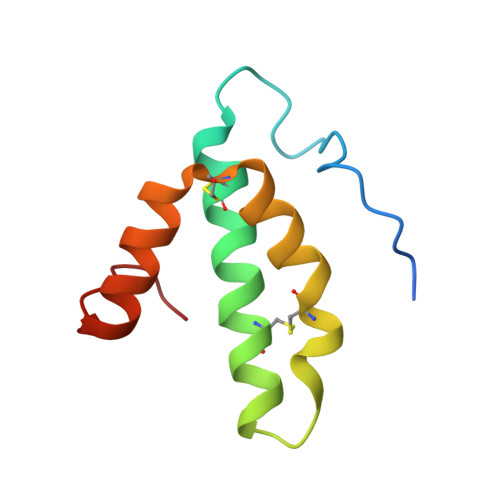
Cytochrome c oxidase (C c O) is a large membrane-bound hemeprotein that catalyzes the reduction of dioxygen to water. Unlike classical dioxygen binding hemeproteins with a heme b group in their active sites, C c O has a unique binuclear center (BNC) composed of a copper atom (Cu B ) and a heme a 3 iron, where O 2 binds and is reduced to water. CO is a versatile O 2 surrogate in ligand binding and escape reactions. Previous time-resolved spectroscopic studies of the CO complexes of bovine C c O (bC c O) revealed that photolyzing CO from the heme a 3 iron leads to a metastable intermediate (Cu B -CO), where CO is bound to Cu B , before it escapes out of the BNC. Here, with a pump-probe based time-resolved serial femtosecond X-ray crystallography, we detected a geminate photoproduct of the bC c O-CO complex, where CO is dissociated from the heme a 3 iron and moved to a temporary binding site midway between the Cu B and the heme a 3 iron, while the locations of the two metal centers and the conformation of Helix-X, housing the proximal histidine ligand of the heme a 3 iron, remain in the CO complex state. This new structure, combined with other reported structures of bC c O, allows for a clearer definition of the ligand dissociation trajectory as well as the associated protein dynamics.
- Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York 10461, United States.
Organizational Affiliation: