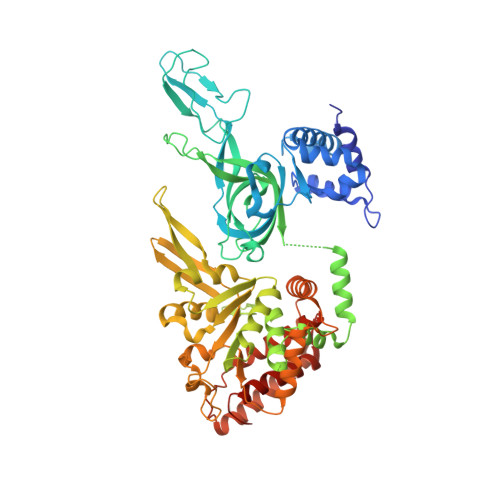
Two Distinct Modes of DNA Binding by an MCM Helicase Enable DNA Translocation.
Meagher, M., Myasnikov, A., Enemark, E.J.(2022) Int J Mol Sci 23
- PubMed: 36499022 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms232314678
- Primary Citation Related Structures:
8EAF, 8EAG, 8EAH, 8EAI, 8EAJ, 8EAK, 8EAL, 8EAM - PubMed Abstract:
A six-subunit ATPase ring forms the central hub of the replication forks in all domains of life. This ring performs a helicase function to separate the two complementary DNA strands to be replicated and drives the replication machinery along the DNA. Disruption of this helicase/ATPase ring is associated with genetic instability and diseases such as cancer. The helicase/ATPase rings of eukaryotes and archaea consist of six minichromosome maintenance (MCM) proteins. Prior structural studies have shown that MCM rings bind one encircled strand of DNA in a spiral staircase, suggesting that the ring pulls this strand of DNA through its central pore in a hand-over-hand mechanism where the subunit at the bottom of the staircase dissociates from DNA and re-binds DNA one step above the staircase. With high-resolution cryo-EM, we show that the MCM ring of the archaeal organism Saccharolobus solfataricus binds an encircled DNA strand in two different modes with different numbers of subunits engaged to DNA, illustrating a plausible mechanism for the alternating steps of DNA dissociation and re-association that occur during DNA translocation.
- Department of Structural Biology, St. Jude Children's Research Hospital, 262 Danny Thomas Place, Memphis, TN 38105, USA.
Organizational Affiliation: