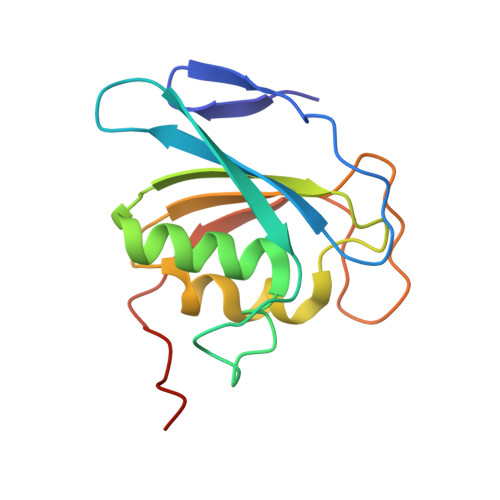
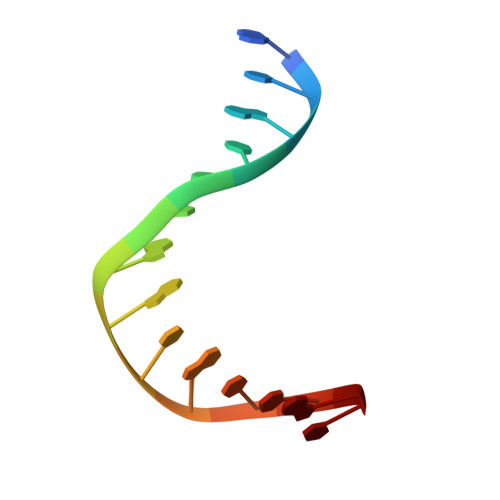
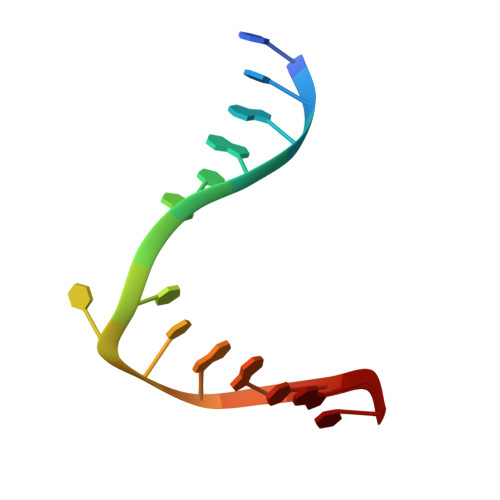
Structural basis of sequence-specific cytosine deamination by double-stranded DNA deaminase toxin DddA.
Yin, L., Shi, K., Aihara, H.(2023) Nat Struct Mol Biol 30: 1153-1159
- PubMed: 37460895 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01034-3
- Primary Citation Related Structures:
8E5D, 8E5E - PubMed Abstract:
The interbacterial deaminase toxin DddA catalyzes cytosine-to-uracil conversion in double-stranded (ds) DNA and enables CRISPR-free mitochondrial base editing, but the molecular mechanisms underlying its unique substrate selectivity have remained elusive. Here, we report crystal structures of DddA bound to a dsDNA substrate containing the 5'-TC target motif. These structures show that DddA binds to the minor groove of a sharply bent dsDNA and engages the target cytosine extruded from the double helix. DddA Phe1375 intercalates in dsDNA and displaces the 5' (-1) thymine, which in turn replaces the target (0) cytosine and forms a noncanonical T-G base pair with the juxtaposed guanine. This tandem displacement mechanism allows DddA to locate a target cytosine without flipping it into the active site. Biochemical experiments demonstrate that DNA base mismatches enhance the DddA deaminase activity and relax its sequence selectivity. On the basis of the structural information, we further identified DddA mutants that exhibit attenuated activity or altered substrate preference. Our studies may help design new tools useful in genome editing or other applications.
- Department of Biochemistry, Molecular Biology, and Biophysics, University of Minnesota, Minneapolis, MN, USA.
Organizational Affiliation: