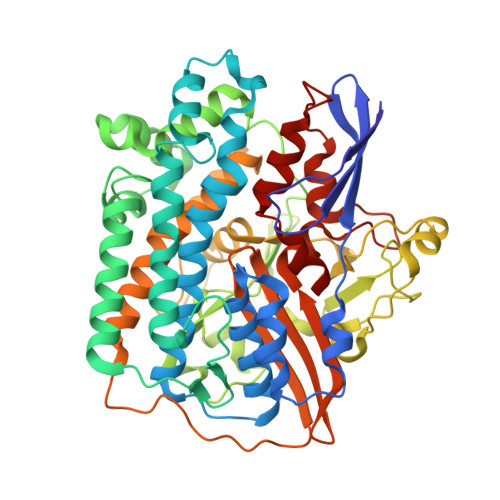
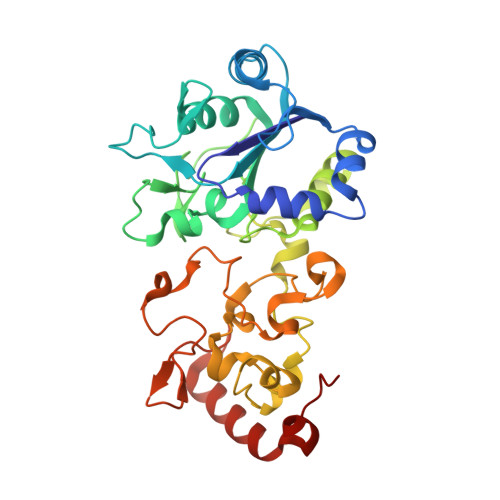
Structural basis for bacterial energy extraction from atmospheric hydrogen.
Grinter, R., Kropp, A., Venugopal, H., Senger, M., Badley, J., Cabotaje, P.R., Jia, R., Duan, Z., Huang, P., Stripp, S.T., Barlow, C.K., Belousoff, M., Shafaat, H.S., Cook, G.M., Schittenhelm, R.B., Vincent, K.A., Khalid, S., Berggren, G., Greening, C.(2023) Nature 615: 541-547
- PubMed: 36890228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-05781-7
- Primary Citation Related Structures:
7UTD, 7UUR, 7UUS, 8DQV - PubMed Abstract:
Diverse aerobic bacteria use atmospheric H 2 as an energy source for growth and survival 1 . This globally significant process regulates the composition of the atmosphere, enhances soil biodiversity and drives primary production in extreme environments 2,3 . Atmospheric H 2 oxidation is attributed to uncharacterized members of the [NiFe] hydrogenase superfamily 4,5 . However, it remains unresolved how these enzymes overcome the extraordinary catalytic challenge of oxidizing picomolar levels of H 2 amid ambient levels of the catalytic poison O 2 and how the derived electrons are transferred to the respiratory chain 1 . Here we determined the cryo-electron microscopy structure of the Mycobacterium smegmatis hydrogenase Huc and investigated its mechanism. Huc is a highly efficient oxygen-insensitive enzyme that couples oxidation of atmospheric H 2 to the hydrogenation of the respiratory electron carrier menaquinone. Huc uses narrow hydrophobic gas channels to selectively bind atmospheric H 2 at the expense of O 2 , and 3 [3Fe-4S] clusters modulate the properties of the enzyme so that atmospheric H 2 oxidation is energetically feasible. The Huc catalytic subunits form an octameric 833 kDa complex around a membrane-associated stalk, which transports and reduces menaquinone 94 Å from the membrane. These findings provide a mechanistic basis for the biogeochemically and ecologically important process of atmospheric H 2 oxidation, uncover a mode of energy coupling dependent on long-range quinone transport, and pave the way for the development of catalysts that oxidize H 2 in ambient air.
- Department of Microbiology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia. rhys.grinter@monash.edu.
Organizational Affiliation: