Stabilization of Cereibacter sphaeroides Photosynthetic Reaction Center by the Introduction of Disulfide Bonds.
Selikhanov, G., Atamas, A., Yukhimchuk, D., Fufina, T., Vasilieva, L., Gabdulkhakov, A.(2023) Membranes (Basel) 13
- PubMed: 36837657 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/membranes13020154
- Primary Citation Related Structures:
8C5X, 8C6K, 8C7C, 8C87, 8C88 - PubMed Abstract:
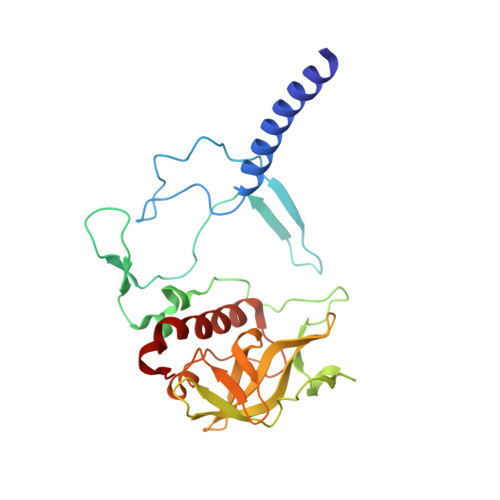
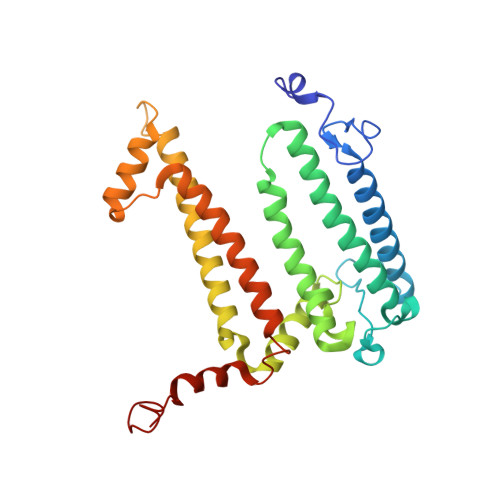
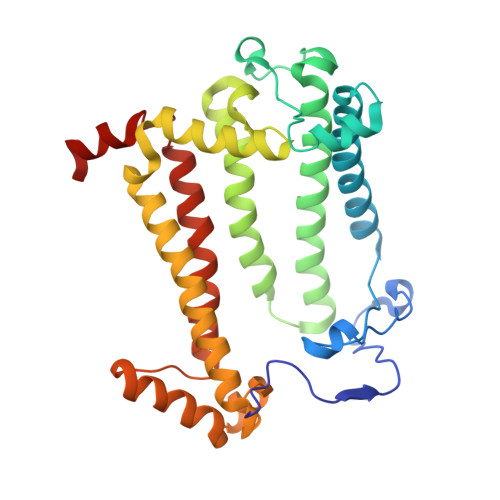
The photosynthetic reaction center of the purple nonsulfur bacterium Cereibacter sphaeroides is a useful model for the study of mechanisms of photoinduced electron transfer and a promising component for photo-bio-electrocatalytic systems. The basic research and technological applications of this membrane pigment-protein complex require effective approaches to increase its structural stability. In this work, a rational design approach to genetically modify the reaction centers by introducing disulfide bonds is used. This resulted in significantly increasing the thermal stability of some of the mutant pigment-protein complexes. The formation of the S-S bonds was confirmed by X-ray crystallography as well as SDS-PAGE, and the optical properties of the reaction centers were studied. The genetically modified reaction centers presented here preserved their ability for photochemical charge separation and could be of interest for basic science and biotechnology.
- Institute of Protein Research, Russian Academy of Sciences, Institutskaya 4, 142290 Pushchino, Moscow Region, Russia.
Organizational Affiliation: