Cryo-EM Helical Structure of dITP-activated KomBC complex
Hao, F., Eddie, T., Bin, W., Min, L.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| KomB, HAM-like protein, Non-canonical purine NTP pyrophosphatase | 184 | Escherichia coli | Mutation(s): 0 Gene Names: GRW56_02065, GRW57_01900 | 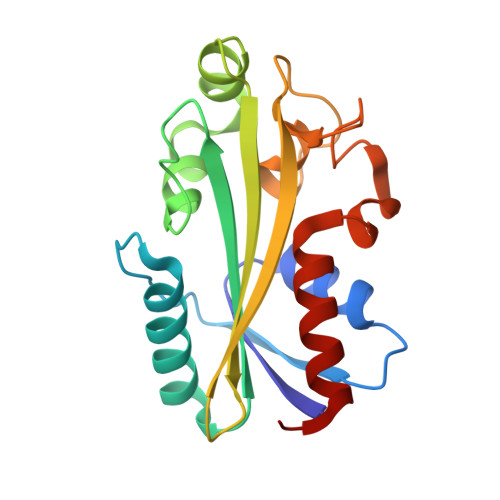 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | K8WC63 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| KomC, SIR2 domain protein, NADase | 264 | Escherichia coli | Mutation(s): 0 Gene Names: GRW56_02060, GRW57_01895 | 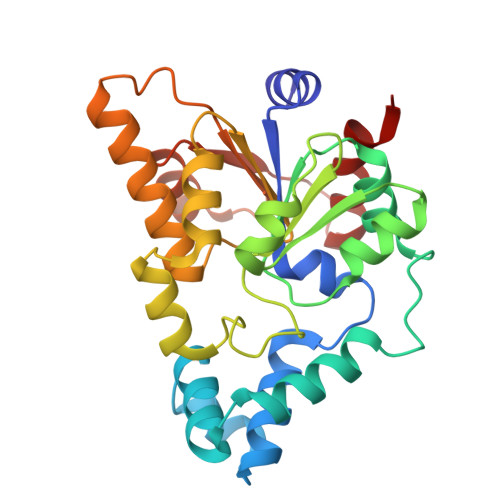 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | K8WJ15 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| Y43 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | AB [auth W] CB [auth U] EB [auth f] GA [auth A] GB [auth Z] | 2'-deoxyinosine 5'-triphosphate C10 H15 N4 O13 P3 UFJPAQSLHAGEBL-RRKCRQDMSA-N |  | ||
| MG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | BB [auth W] DB [auth U] FB [auth f] HA [auth A] HB [auth Z] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |