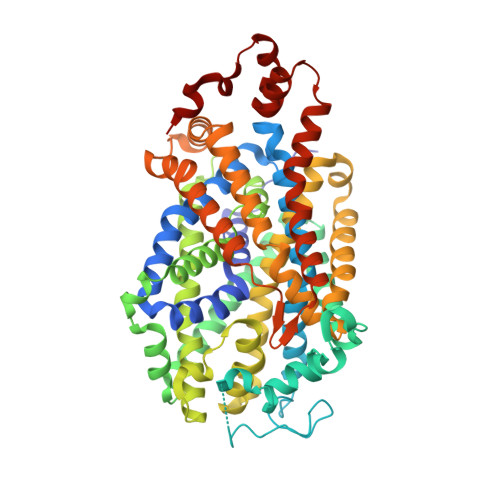
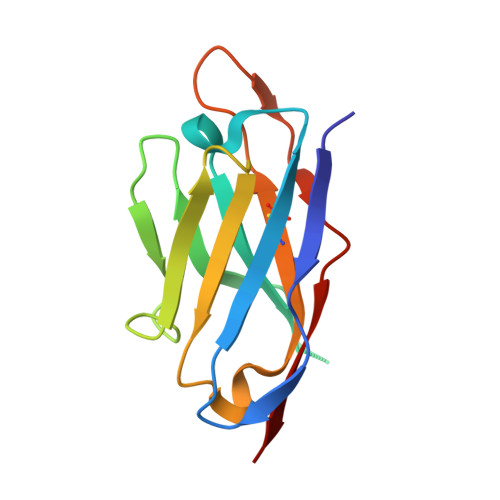
Mechanistic insights of substrate transport and inhibitor binding revealed by high-resolution structures of human norepinephrine transporter.
Song, A., Wu, X.(2024) Cell Res 34: 810-813
- PubMed: 39223283 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-024-01024-0
- Primary Citation Related Structures:
8ZOY, 8ZP1, 8ZP2, 8ZPB - Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou, Zhejiang, China.
Organizational Affiliation: