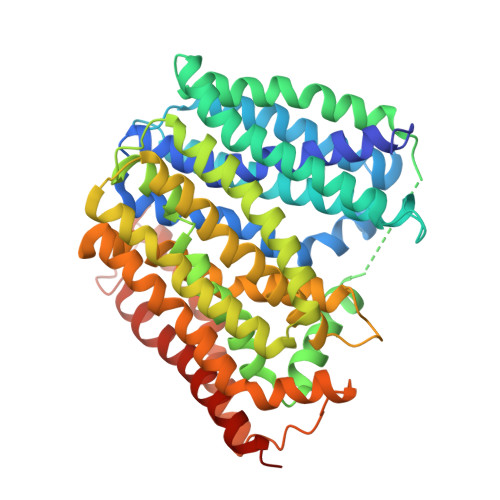
Structural and functional insights of AmpG in muropeptide transport and multiple beta-lactam antibiotics resistance.
Chang, N., Kim, H., Kim, U., Cho, Y., Yoo, Y., Lee, H., Kim, J.W., Kim, M.S., Lee, J., Cho, Y.L., Kim, K., Yong, D., Cho, H.S.(2025) Nat Commun 16: 5744-5744
- PubMed: 40593790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-61169-3
- Primary Citation Related Structures:
8ZBB, 8ZGZ, 8ZKE, 9J9Z - PubMed Abstract:
Anhydromuropeptide permease (AmpG) is a transporter protein located in the inner membrane of certain gram -negative bacteria, involved in peptidoglycan (PG) recycling and β-lactamase induction. Decreased AmpG function reduces resistance of antibiotic-resistant bacteria to β-lactam antibiotics. Therefore, AmpG-targeting inhibitors are promising 'antibiotic adjuvants'. However, as the tertiary structure of AmpG has not yet been identified, the development of targeted inhibitors remains challenging. We present four cryo-electron microscopy (cryo-EM) structures: the apo-inward and apo-outward state structures and the inward-occluded and outward states complexed with the substrate GlcNAc-1,6-anhMurNAc. Through functional analysis and molecular dynamics (MD) simulations, we identified motif A, which stabilizes the outward state, substrate-binding pocket, and protonation-related residues. Based on the structure of AmpG and our experimental results, we propose a muropeptide transport mechanism for AmpG. A deeper understanding of its structure and transport mechanism provides a foundation for the development of antibiotic adjuvants.
- Department of Systems Biology and Division of Life Sciences, Yonsei University, 50 Yonsei-ro, Seoul, Republic of Korea.
Organizational Affiliation: