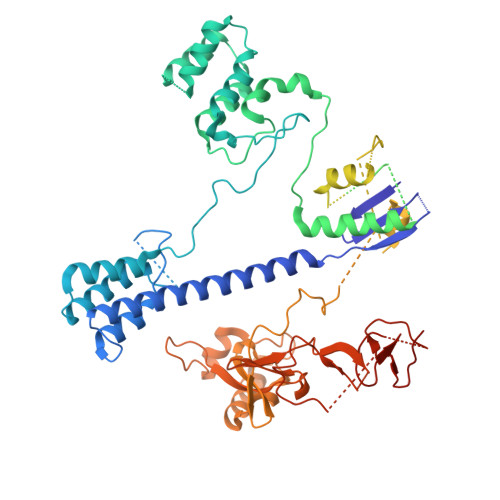
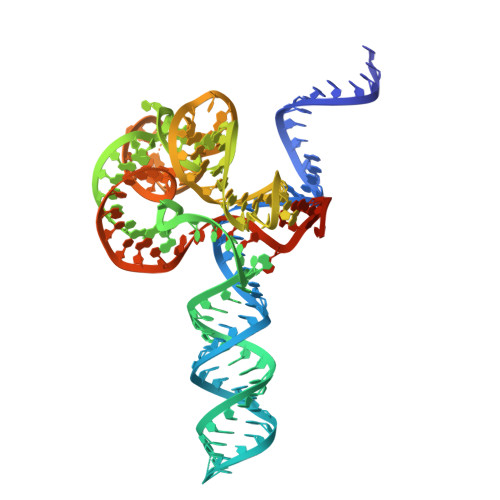
Insights into the compact CRISPR-Cas9d system.
Yang, J., Wang, T., Huang, Y., Long, Z., Li, X., Zhang, S., Zhang, L., Liu, Z., Zhang, Q., Sun, H., Zhang, M., Yin, H., Liu, Z., Zhang, H.(2025) Nat Commun 16: 2462-2462
- PubMed: 40075056 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-57455-9
- Primary Citation Related Structures:
8ZDR, 8ZQ9 - PubMed Abstract:
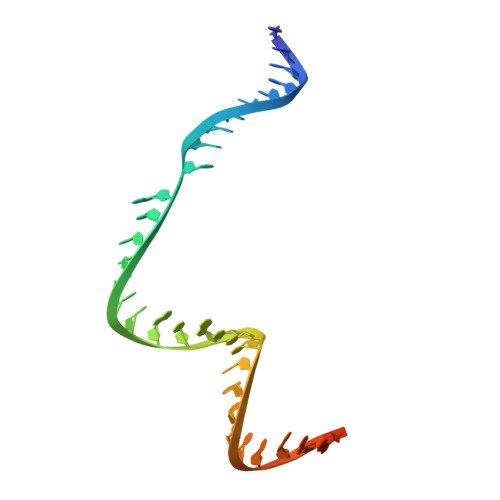
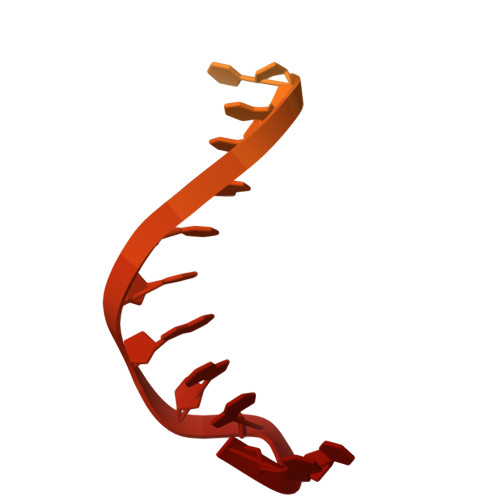
Cas9d, the smallest known member of the Cas9 family, employs a compact domain architecture for effective target cleavage. However, the underlying mechanism remains unclear. Here, we present the cryo-EM structures of the Cas9d-sgRNA complex in both target-free and target-bound states. Biochemical assays elucidated the PAM recognition and DNA cleavage mechanisms of Cas9d. Structural comparisons revealed that at least 17 base pairs in the guide-target heteroduplex is required for nuclease activity. Beyond its typical role as an adaptor between Cas9 enzymes and targets, the sgRNA also provides structural support and functional regulation for Cas9d. A segment of the sgRNA scaffold interacts with the REC domain to form a functional target recognition module. Upon target binding, this module undergoes a coordinated conformational rearrangement, enabling heteroduplex propagation and facilitating nuclease activity. This hybrid functional module precisely monitors heteroduplex complementarity, resulting in a lower mismatch tolerance compared to SpyCas9. Moreover, structure-guided engineering in both the sgRNA and Cas9d protein led to a more compact Cas9 system with well-maintained nuclease activity. Altogether, our findings provide insights into the target recognition and cleavage mechanisms of Cas9d and shed light on the development of high-fidelity mini-CRISPR tools.
- State Key Laboratory of Experimental Hematology, Tianjin Institute of Immunology, The Province and Ministry Co-sponsored Collaborative Innovation Center for Medical Epigenetics, Key Laboratory of Immune Microenvironment and Disease (Ministry of Education), School of Basic Medical Sciences, Tianjin Medical University, Tianjin, China.
Organizational Affiliation: