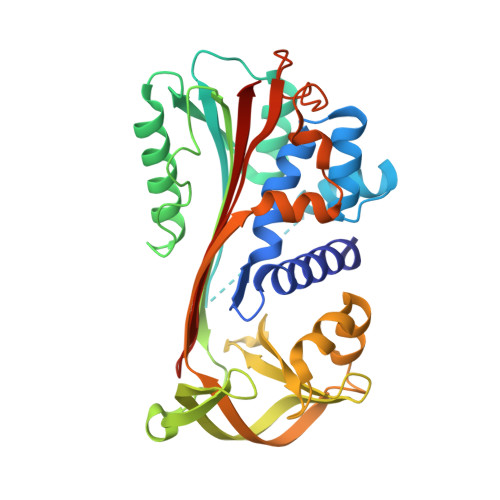
Crystallization and crystallographic studies of human serine protease inhibitor (serpin) B9.
Yan, T., Zhou, A.(2024) Acta Crystallogr F Struct Biol Commun 80: 286-293
- PubMed: 39382088 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X24009439
- Primary Citation Related Structures:
8ZCR - PubMed Abstract:
Serine protease inhibitor B9 (serpin B9, also known as protease inhibitor 9 or PI9) plays a critical role in regulating the immune response by specifically inhibiting granzyme B, a serine protease found in cytotoxic T lymphocytes and natural killer cells. Despite its potential as an anticancer drug target, the structural details of serpin B9 have remained elusive until now. In this study, a cleaved form of recombinant human serpin B9 was successfully prepared and crystallized. The crystals belonged to space group P2 1 2 1 2 1 , with unit-cell parameters a = 68.51, b = 82.32, c = 101.17 Å, and an X-ray diffraction data set was collected at 1.9 Å resolution. The structure shows that serpin B9 adopts a relaxed conformation, with its cleaved reactive-centre loop inserted into the central β-sheet. Unlike other serpins, serpin B9 shows significant structural deviations around helix D, with a larger surface cavity, which could serve as a promising target for small-molecule inhibitors.
- Department of Pathophysiology, Shanghai Jiaotong University School of Medicine, 280 South Chongqing Road, Shanghai 200025, People's Republic of China.
Organizational Affiliation: