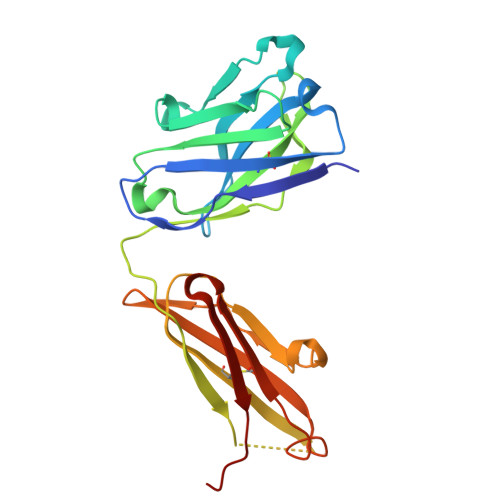
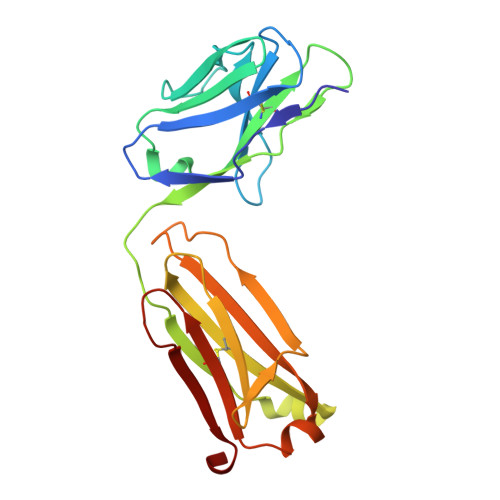
Cryo-EM structure of the bacterial intramembrane metalloprotease RseP in the substrate-bound state.
Asahi, K., Hirose, M., Aruga, R., Shimizu, Y., Tajiri, M., Tanaka, T., Adachi, Y., Tanaka, Y., Kaneko, M.K., Kato, Y., Akashi, S., Akiyama, Y., Hizukuri, Y., Kato, T., Nogi, T.(2025) Sci Adv 11: eadu0925-eadu0925
- PubMed: 40009668 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adu0925
- Primary Citation Related Structures:
8ZAY, 9J82, 9J83 - PubMed Abstract:
Site-2 proteases (S2Ps), conserved intramembrane metalloproteases that maintain cellular homeostasis, are associated with chronic infection and persistence leading to multidrug resistance in bacterial pathogens. A structural model of how S2Ps discriminate and accommodate substrates could help us develop selective antimicrobial agents. We previously proposed that the Escherichia coli S2P RseP unwinds helical substrate segments before cleavage, but the mechanism for accommodating a full-length membrane-spanning substrate remained unclear. Our present cryo-EM analysis of Aquifex aeolicus RseP ( Aa RseP) revealed that a substrate-like membrane protein fragment from the expression host occupied the active site while spanning a transmembrane cavity that is inaccessible via lateral diffusion. Furthermore, in vivo photocrosslinking supported that this substrate accommodation mode is recapitulated on the cell membrane. Our results suggest that the substrate accommodation by threading through a conserved membrane-associated region stabilizes the substrate-complex and contributes to substrate discrimination on the membrane.
- Graduate School of Medical Life Science, Yokohama City University, 1-7-29 Suehiro-cho, Tsurumi-ku, Yokohama 230-0045, Japan.
Organizational Affiliation: