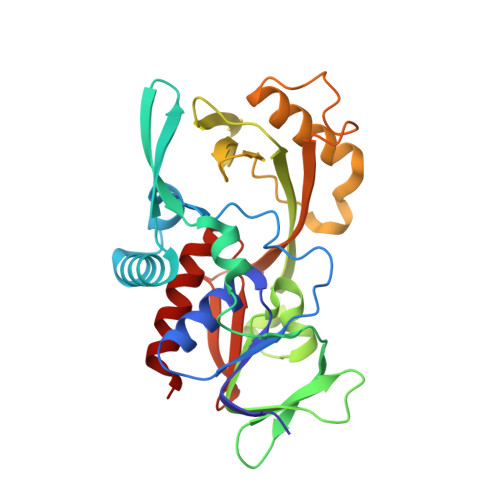
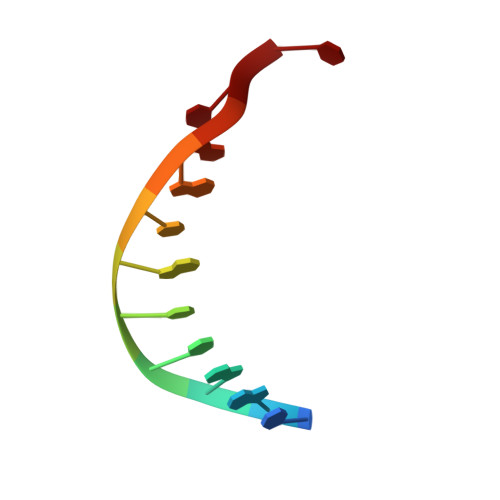
Structural insights into the biosynthetic mechanism of N alpha-GlyT and 5-NmdU hypermodifications of DNA.
Wen, Y., Guo, W., Meng, C., Yang, J., Xu, S., Chen, H., Gan, J., Wu, B.(2024) Nucleic Acids Res 52: 11083-11097
- PubMed: 39268585 Search on PubMed
- DOI: https://doi.org/10.1093/nar/gkae784
- Primary Citation Related Structures:
8Z2M, 8Z2N, 8Z2O, 8Z79 - PubMed Abstract:
DNA hypermodifications are effective weapons for phages to cope with the defense system of bacteria. The biogenesis of DNA hypermodification in phages involves multiple steps, from the modified deoxynucleotide monophosphates to the final hypermodification on the DNA chains. PseudomonasPaMx11 gp46 and gp47 encode the enzymes for sequentially converting 5-phosphomethyl-2'-deoxyuridine to 5-Nα-glycinylthymidine and 5-aminomethyl-2'-deoxyuridine. Here, we have determined the crystal structures of gp46 and gp47 in their apo and double-stranded DNA (dsDNA)-bound forms. We uncovered their dsDNA recognition properties and identified the critical residues for the catalytic reactions. Combined with in vitro biochemical studies, we proposed a plausible reaction scheme for gp46 and gp47 in converting these DNA hypermodifications. Our studies will provide the structural basis for future bioengineering of the synthetic pathway of hypermodification and identifying new modifications in mammals by enzyme-assisted sequencing methods.
- Guangdong Provincial Key Laboratory of Malignant Tumor Epigenetics and Gene Regulation, Guangdong-Hong Kong Joint Laboratory for RNA Medicine, Medical Research Center, Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University, 107 Yanjiang West Road, Guangzhou 510120, China.
Organizational Affiliation: