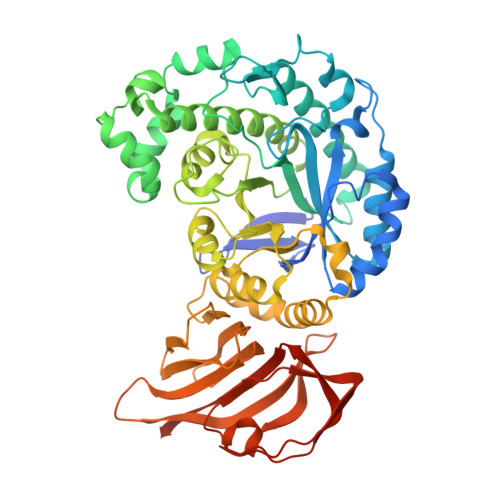
Structure and function of a beta-1,2-galactosidase from Bacteroides xylanisolvens, an intestinal bacterium.
Nakazawa, Y., Kageyama, M., Matsuzawa, T., Liang, Z., Kobayashi, K., Shimizu, H., Maeda, K., Masuhiro, M., Motouchi, S., Kumano, S., Tanaka, N., Kuramochi, K., Nakai, H., Taguchi, H., Nakajima, M.(2025) Commun Biol 8: 66-66
- PubMed: 39820076 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-025-07494-1
- Primary Citation Related Structures:
8Z43, 8Z47, 8Z48 - PubMed Abstract:
Galactosides are major carbohydrates that are found in plant cell walls and various prebiotic oligosaccharides. Studying the detailed biochemical functions of β-galactosidases in degrading these carbohydrates is important. In particular, identifying β-galactosidases with new substrate specificities could help in the production of potentially beneficial oligosaccharides. In this study, we identify a β-galactosidase with novel substrate specificity from Bacteroides xylanisolvens, an intestinal bacterium. The enzyme do not show hydrolytic activity toward natural β-galactosides during the first screening. However, when α-D-galactosyl fluoride (α-GalF) as a donor substrate and galactose or D-fucose as an acceptor substrate are incubated with a nucleophile mutant, reaction products are detected. The galactobiose produced from the α-GalF and galactose is identified as β-1,2-galactobiose using NMR. Kinetic analysis reveals that this enzyme effectively hydrolyzes β-1,2-galactobiose and β-1,2-galactotriose. In the complex structure with methyl β-galactopyranose as a ligand, the ligand is only located at subsite +1. The 2-hydroxy group and the anomeric methyl group of methyl β-galactopyranose faces in the direction of subsite -1 and the solvent, respectively. This observation is consistent with the substrate specificity of the enzyme regarding linkage position and chain length. Overall, we conclude that the enzyme is a β-galactosidase acting on β-1,2-galactooligosaccharides.
- Department of Applied Biological Science, Faculty of Science and Technology, Tokyo University of Science, 2641 Yamazaki, Noda, Chiba, 278-8510, Japan.
Organizational Affiliation: