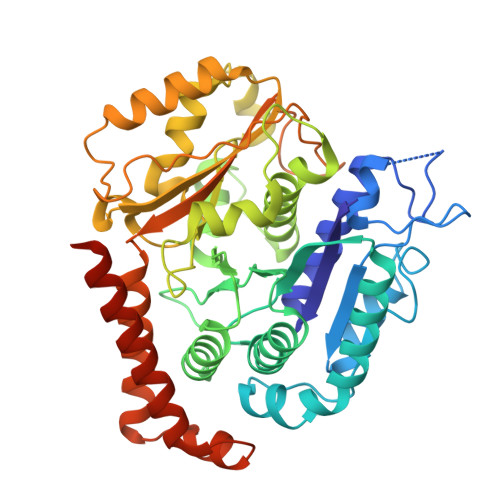
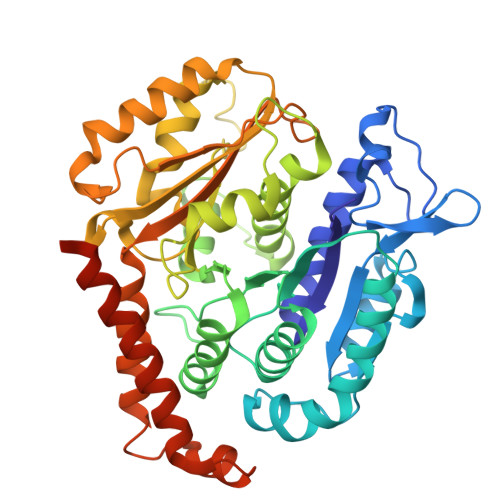
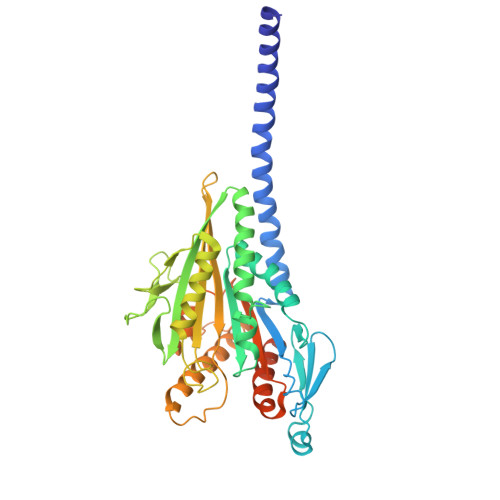
Structural analysis of a motor with increased mechanical output reveals new transitions in kinesin microtubule motility.
Shibata, S., Wang, M.Y., Imasaki, T., Shigematsu, H., Ugarte La Torre, D., Wei, Y., Jobichen, C., Hagio, H., Sivaraman, J., Sugita, Y., Endow, S.A., Nitta, R.(2026) Sci Rep 16: 487-487
- PubMed: 41491787 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-025-28573-7
- Primary Citation Related Structures:
8YUE, 8YY2, 8YY3, 8YY4, 8YY5 - PubMed Abstract:
Kinesin motors use ATP to produce force in cells, yet the conformational changes that generate force remain uncertain. Here, we report structural and mechanistic insights into a minus-end-directed kinesin-14 that exhibits increased mechanical output – the variant motor binds microtubules more tightly and moves with faster velocity than wild type. High-resolution structures, together with molecular dynamics simulations, reveal previously unobserved transitions in the nucleotide hydrolysis cycle. ADP release, triggered by microtubule binding, is coupled to twisting of the central β-sheet and stabilization of the stalk prior to the power stroke. ATP binding induces stalk fluctuations and a swing of the neck mimic, an element analogous to the kinesin-1 neck linker, resembling neck linker docking in plus-end-directed kinesins. The power stroke, characterized by a large stalk rotation, is followed by motor detachment from microtubules. The subsequent recovery stroke occurs while the motor is bound to ADP and free Pi, accompanied by β-strand-to-loop transitions, or β-sheet melting, implying that β-sheet refolding facilitates Pi release. The observed twisting and melting identify the central β-sheet as the long-sought elastic element or spring required for motor force production. The transitions we observe in kinesin-14 may also apply to other kinesins – this remains to be tested. The online version contains supplementary material available at 10.1038/s41598-025-28573-7.
- Division of Structural Medicine and Anatomy, Department of Physiology and Cell Biology, Kobe University Graduate School of Medicine, 650-0017, Kobe, Japan.
Organizational Affiliation: