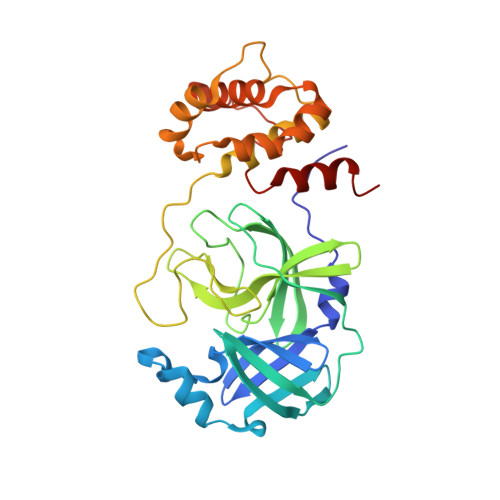
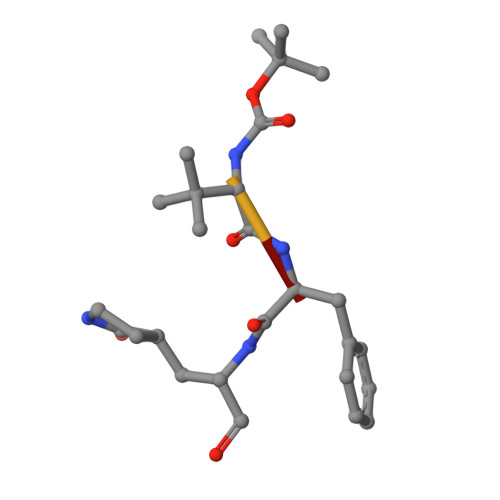
Discovery, Biological Activity, and Structural Mechanism of a Potent Inhibitor of SARS-CoV-2 Main Protease
Wang, J., Sang, X.H., Zheng, W.Y., Chan, J.F., Zhou, J., Xu, Y., Fu, L.F., Feng, Y., Han, P., Tsang, J.O., Yuan, S.F., An, J., Yuen, K.Y., Qi, J.X., Huang, Z.W.To be published.