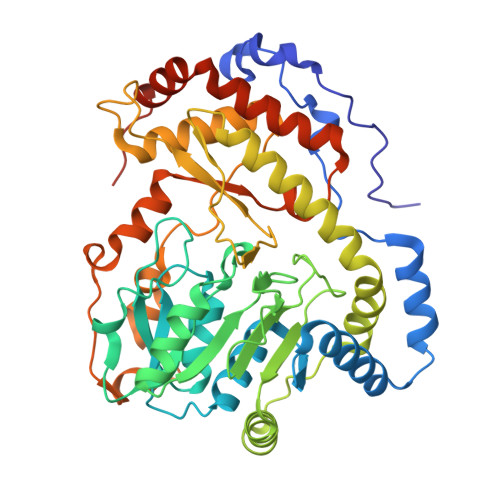
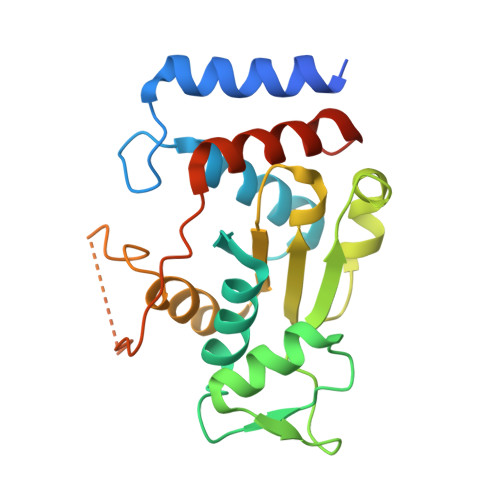
Crystal structure of a novel heterooligomeric aminotransferase from Serratia sp. ATCC 39006 provides insights into function.
Pramono, H., Yoshida, A., Hirashima, Y., Sone, Y., Terada, T., Kosono, S., Nishiyama, M.(2025) FEBS Lett 599: 74-88
- PubMed: 39618122 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.15068
- Primary Citation Related Structures:
8Y96, 8Y97, 8Y98 - PubMed Abstract:
Serratia sp. ATCC 39006 has two tandemly positioned genes, ser4 and ser5, both annotated as sugar aminotransferases, in a putative secondary metabolite biosynthetic gene cluster. Ser5 possesses a complete fold-type I aminotransferase fold, while Ser4 lacks the N- and C-terminal regions and a catalytically important lysine residue of fold-type I aminotransferase. We herein revealed that Ser4 and Ser5 formed a heterotetrameric complex (SerTA) with aminotransferase activity and determined the crystal structures. MD simulations and activity assays with SerTA variants indicated that residues from helix α-8* of inactive Ser4 are important for activity, confirming the importance of heterocomplex formation for activity. Furthermore, the structures suggest that SerTA recognizes a substrate loaded on the carrier protein.
- Graduate School of Agriculture and Life Sciences, The University of Tokyo, Bunkyo-ku, Japan.
Organizational Affiliation: