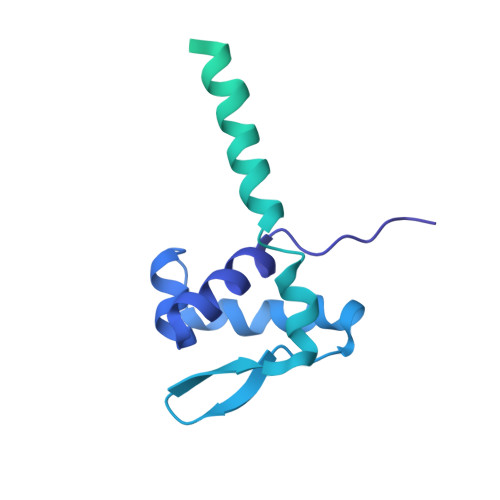
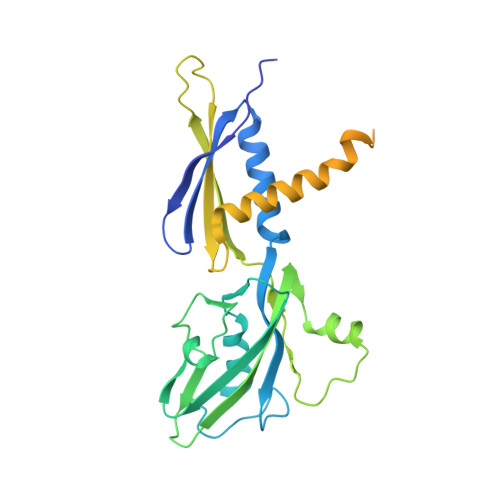
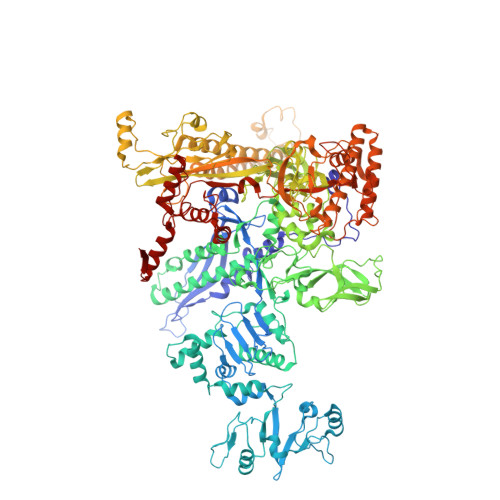
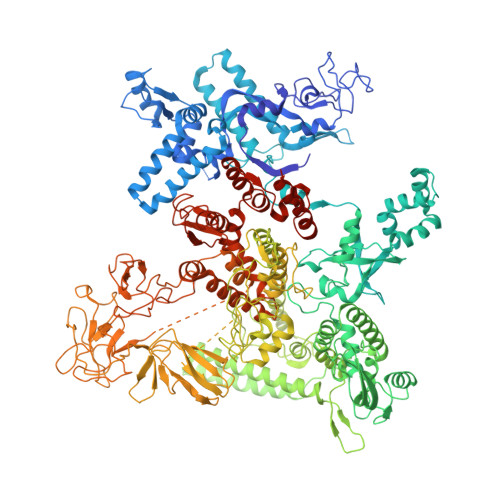
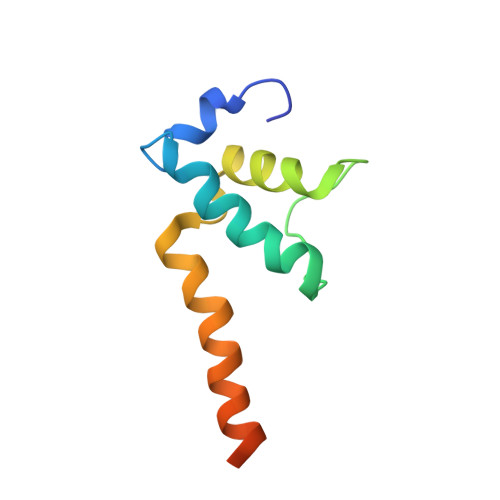
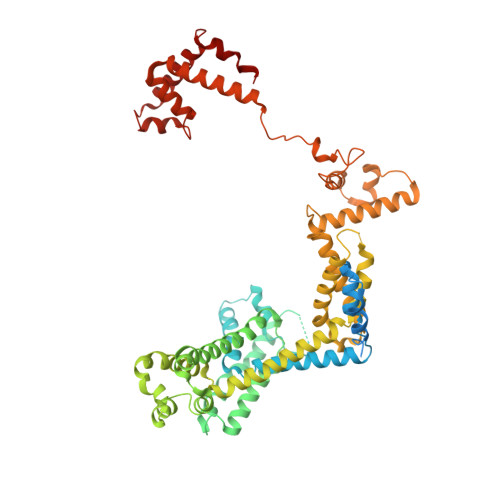
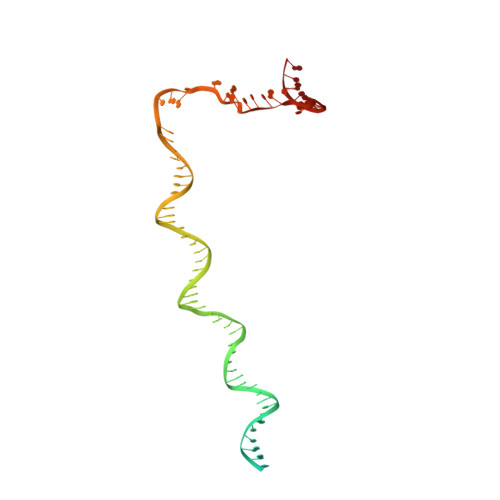
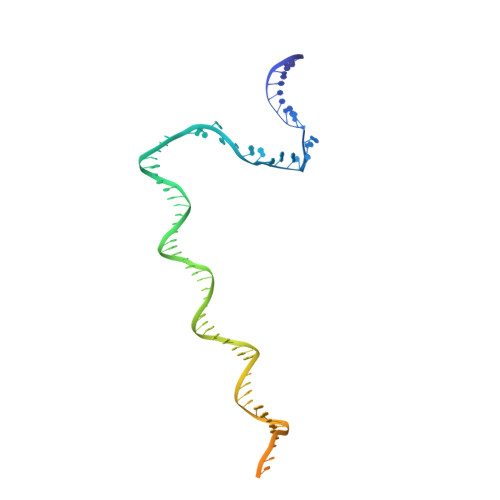
NMR analysis of a loop-bulge structure of UUCGA pentaloop.
Ichijo, R., Kawai, G.(2024) Biochem Biophys Res Commun 691: 149327-149327
- PubMed: 38039839 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2023.149327
- Primary Citation Related Structures:
8K8B, 8Y6U - PubMed Abstract:
Although structures of many RNA loops, such as GNRA and UNCG tetraloops, were well known, it is still possible to find more RNA structures. In the present study, solution structure of an RNA fragment having UUCGA pentaloop was analyzed by NMR spectroscopy. It was found that the UUCG tetraloop is formed and the adenosine residue at the 3' side of the tetraloop is bulged out. The characteristic motif of the loop-bulge structure has also been found in other RNAs including CUUGU and CUGGC pentaloops. Along with the recently found T-hairpin structure with a UUUGAUU loop, in which UUUGA pentaloop and UU bulge are formed, the loop-bulge structures can be categorized as an RNA motif and it may be called as the integrated structure loop, I-loop.
- Graduate school of Advanced Engineering, Chiba Institute of Technology, Tsudanuma, Narashino, Chiba, 275-0016, Japan.
Organizational Affiliation: