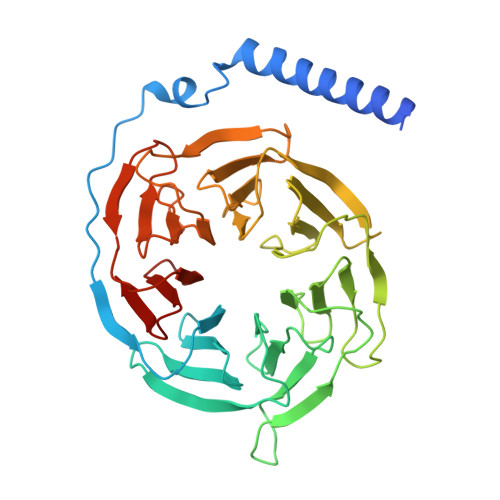
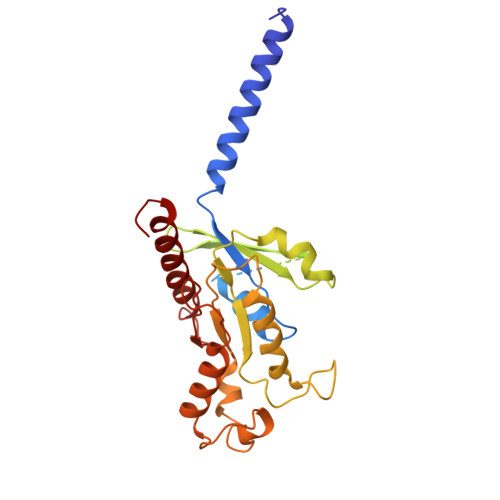
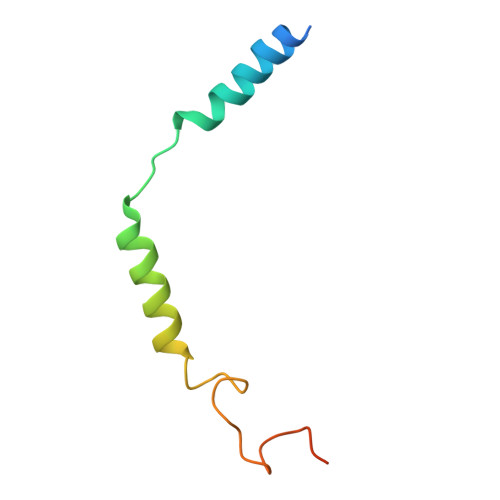
Cryo-EM structure of small-molecule agonist bound delta opioid receptor-G i complex enables discovery of biased compound.
Cheng, L., Miao, Z., Liu, S., Li, Z., Fu, H., Xu, C., Hu, S., Zhao, C., Liu, Y., Zhao, T., Liu, W., Wang, H., Liu, R., Yan, W., Tang, X., Liu, J., Shao, Z., Ke, B.(2024) Nat Commun 15: 8284-8284
- PubMed: 39333070 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-52601-1
- Primary Citation Related Structures:
8Y45 - PubMed Abstract:
Delta opioid receptor (δOR) plays a pivotal role in modulating human sensation and emotion. It is an attractive target for drug discovery since, unlike Mu opioid receptor, it is associated with low risk of drug dependence. Despite its potential applications, the pharmacological properties of δOR, including the mechanisms of activation by small-molecule agonists and the complex signaling pathways it engages, as well as their relation to the potential side effects, remain poorly understood. In this study, we use cryo-electron microscopy (cryo-EM) to determine the structure of the δOR-G i complex when bound to a small-molecule agonist (ADL5859). Moreover, we design a series of probes to examine the key receptor-ligand interaction site and identify a region involved in signaling bias. Using ADL06 as a chemical tool, we elucidate the relationship between the β-arrestin pathway of the δOR and its biological functions, such as analgesic tolerance and convulsion activities. Notably, we discover that the β-arrestin recruitment of δOR might be linked to reduced gastrointestinal motility. These insights enhance our understanding of δOR's structure, signaling pathways, and biological functions, paving the way for the structure-based drug discovery.
- Department of Anesthesiology, Laboratory of Anesthesia and Critical Care Medicine, National-Local Joint Engineering Research Centre of Translational Medicine of Anesthesiology, State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, Sichuan, China.
Organizational Affiliation: