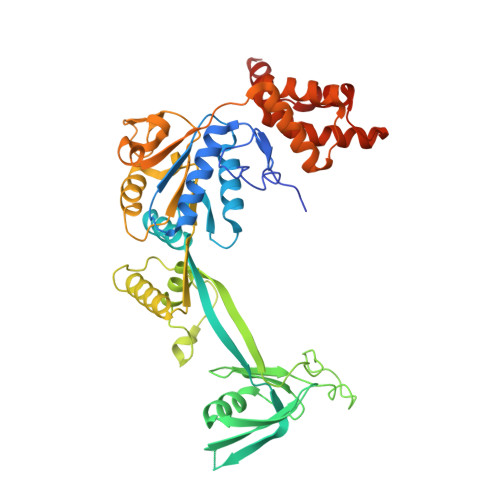
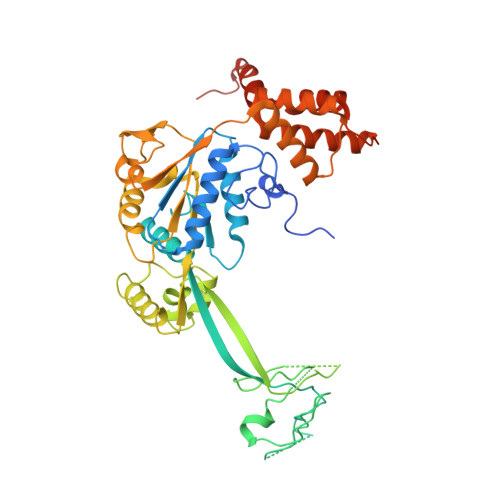
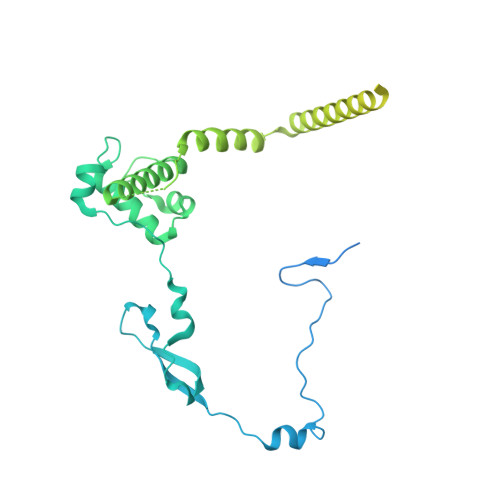
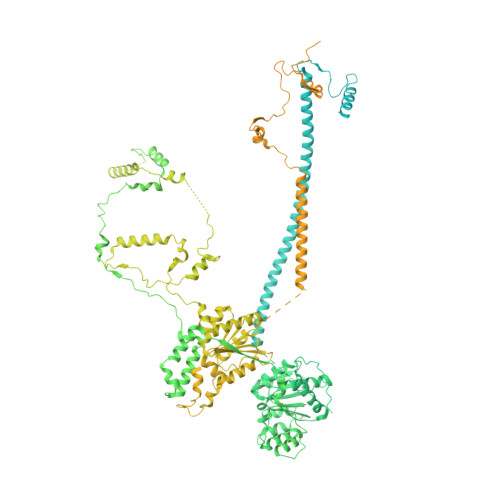
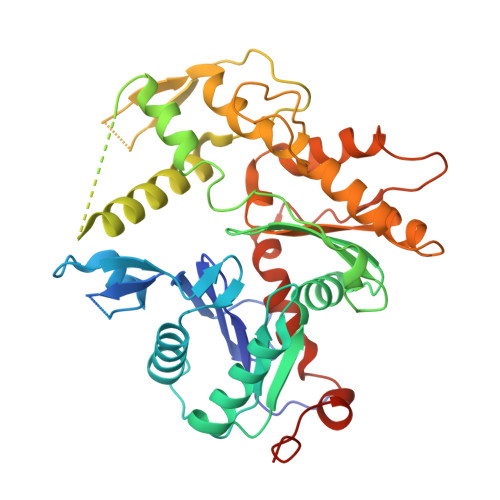
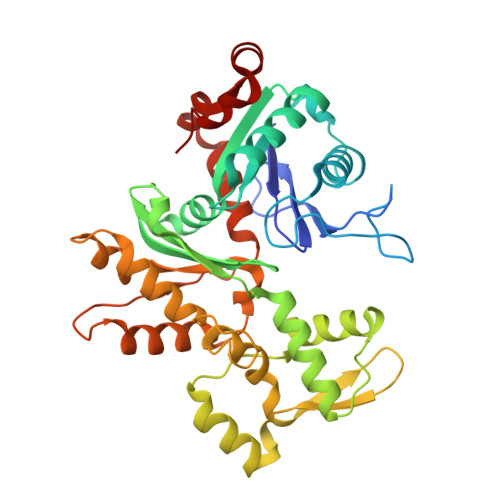
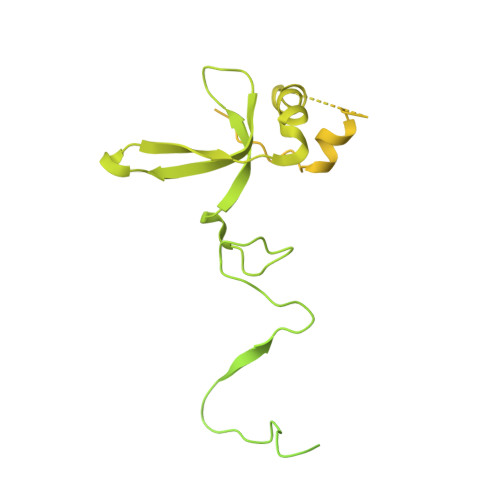
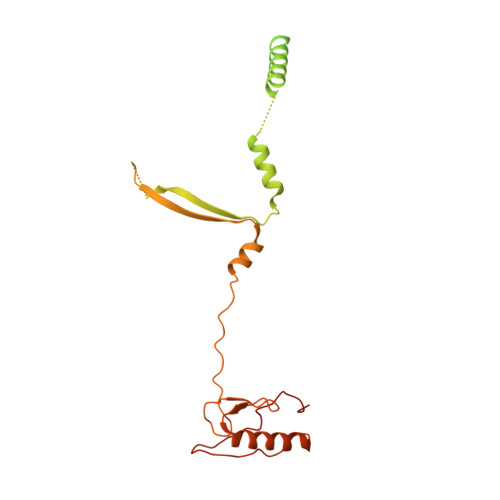
Structure of the human TIP60 complex.
Chen, K., Wang, L., Yu, Z., Yu, J., Ren, Y., Wang, Q., Xu, Y.(2024) Nat Commun 15: 7092-7092
- PubMed: 39154037 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-51259-z
- Primary Citation Related Structures:
8XVG, 8XVT, 8XVV - PubMed Abstract:
Mammalian TIP60 is a multi-functional enzyme with histone acetylation and histone dimer exchange activities. It plays roles in diverse cellular processes including transcription, DNA repair, cell cycle control, and embryonic development. Here we report the cryo-electron microscopy structures of the human TIP60 complex with the core subcomplex and TRRAP module refined to 3.2-Å resolution. The structures show that EP400 acts as a backbone integrating the motor module, the ARP module, and the TRRAP module. The RUVBL1-RUVBL2 hexamer serves as a rigid core for the assembly of EP400 ATPase and YL1 in the motor module. In the ARP module, an ACTL6A-ACTB heterodimer and an extra ACTL6A make hydrophobic contacts with EP400 HSA helix, buttressed by network interactions among DMAP1, EPC1, and EP400. The ARP module stably associates with the motor module but is flexibly tethered to the TRRAP module, exhibiting a unique feature of human TIP60. The architecture of the nucleosome-bound human TIP60 reveals an unengaged nucleosome that is located between the core subcomplex and the TRRAP module. Our work illustrates the molecular architecture of human TIP60 and provides architectural insights into how this complex is bound by the nucleosome.
- Fudan University Shanghai Cancer Center, Institutes of Biomedical Sciences, State Key Laboratory of Genetic Engineering and Shanghai Key Laboratory of Medical Epigenetics, Shanghai Medical College of Fudan University, Shanghai, 200032, China.
Organizational Affiliation: