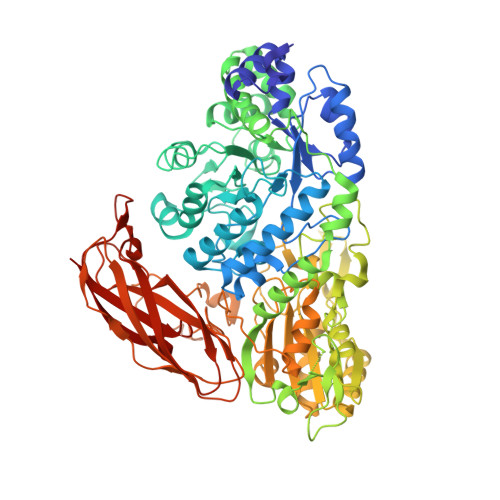
A trapped covalent intermediate as a key catalytic element in the hydrolysis of a GH3 beta-glucosidase: An X-ray crystallographic and biochemical study.
Hu, C., Wang, Y., Wang, W., Cui, W., Jia, X., Mayo, K.H., Zhou, Y., Su, J., Yuan, Y.(2024) Int J Biol Macromol 265: 131131-131131
- PubMed: 38527679 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2024.131131
- Primary Citation Related Structures:
8XRT, 8XRU, 8XRV, 8XRX - PubMed Abstract:
Glycoside hydrolases (GHs) are industrially important enzymes that hydrolyze glycosidic bonds in glycoconjugates. In this study, we found a GH3 β-glucosidase (CcBgl3B) from Cellulosimicrobium cellulans sp. 21 was able to selectively hydrolyze the β-1,6-glucosidic bond linked glucose of ginsenosides. X-ray crystallographic studies of the ligand complex ginsenoside-specific β-glucosidase provided a novel finding that support the catalytic mechanism of GH3. The substrate was clearly identified within the catalytic center of wild-type CcBgl3B, revealing that the C1 atom of the glucose was covalently bound to the Oδ1 group of the conserved catalytic nucleophile Asp264 as an enzyme-glycosyl intermediate. The glycosylated Asp264 could be identified by mass spectrometry. Through site-directed mutagenesis studies with Asp264, it was found that the covalent intermediate state formed by Asp264 and the substrate was critical for catalysis. In addition, Glu525 variants (E525A, E525Q and E525D) showed no or marginal activity against pNPβGlc; thus, this residue could supply a proton for the reaction. Overall, our study provides an insight into the catalytic mechanism of the GH3 enzyme CcBgl3B.
- Engineering Research Center of Glycoconjugates Ministry of Education, Jilin Provincial Key Laboratory of Chemistry and Biology of Changbai Mountain Natural Drugs, School of Life Sciences, Northeast Normal University, Changchun 130024, China.
Organizational Affiliation: