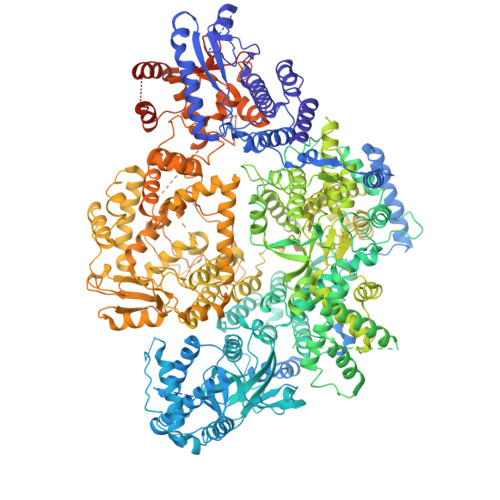
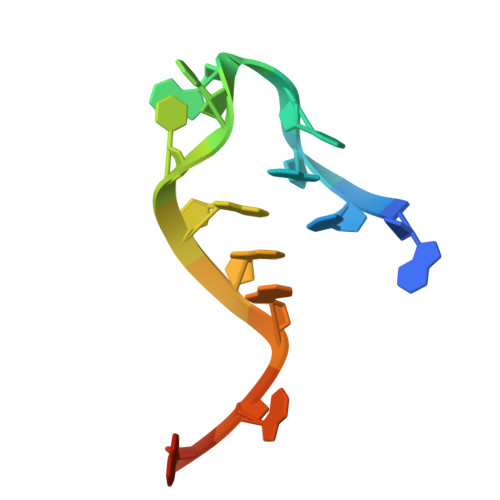
An adenosine analog shows high antiviral potency against coronavirus and arenavirus mainly through an unusual base pairing mode.
Jia, X., Jing, X., Li, M., Gao, M., Zhong, Y., Li, E., Liu, Y., Li, R., Yao, G., Liu, Q., Zhou, M., Hou, Y., An, L., Hong, Y., Li, S., Zhang, J., Wang, W., Zhang, K., Gong, P., Chiu, S.(2024) Nat Commun 15: 10750-10750
- PubMed: 39737930 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-54918-3
- Primary Citation Related Structures:
8XKO, 8XPO, 8XPP - PubMed Abstract:
By targeting the essential viral RNA-dependent RNA polymerase (RdRP), nucleoside analogs (NAs) have exhibited great potential in antiviral therapy for RNA virus-related diseases. However, most ribose-modified NAs do not present broad-spectrum features, likely due to differences in ribose-RdRP interactions across virus families. Here, we show that HNC-1664, an adenosine analog with modifications both in ribose and base, has broad-spectrum antiviral activity against positive-strand coronaviruses and negative-strand arenaviruses. Importantly, treatment with HNC-1664 demonstrate anti-SARS-CoV-2 efficacy in infected K18-human ACE2 mice, with reduced viral titer and mortality, as well as improved lung injury. Enzymology data demonstrate that HNC-1664 inhibits RNA synthesis mainly at the pre-catalysis stage. The cryo-EM structures of HNC-1664-bound RdRP-RNA complexes from both SARS-CoV-2 and LASV reveal an unusual base pairing mode of HNC-1664 in part due to its base modification, thus revealing its great potency in binding but not catalysis. Under certain circumstances, 1664-TP can be slowly incorporated by RdRP through regular Watson-Crick base pairing, as evidenced by enzymology data and an HNC-1664-incorporated crystal structure of the RdRP-RNA complex. Overall, HNC-1664 achieves broad-spectrum characteristics by favoring an alternative base pairing strategy to non-catalytically block RNA synthesis, providing a novel concept for the rational development of NA drugs.
- Key Laboratory of Special Pathogens and Biosafety, Wuhan Institute of Virology, Center for Biosafety Mega-Science, Chinese Academy of Sciences, No. 262 Jin Long Street, Wuhan, Hubei, China.
Organizational Affiliation: