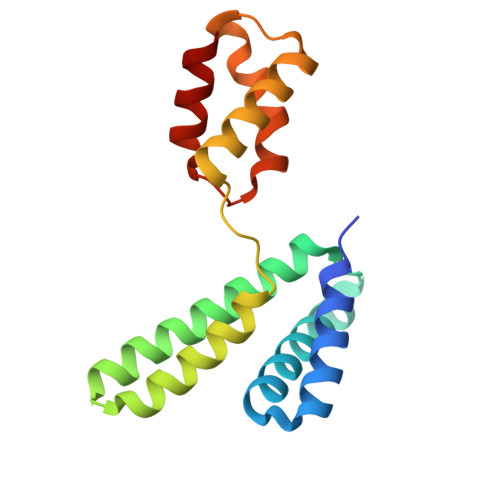
Structural analysis of the YqeY proteins from Campylobacter jejuni and Vibrio parahaemolyticus.
Kim, S.Y., Yoon, S.I.(2024) Biochem Biophys Res Commun 695: 149485-149485
- PubMed: 38211535 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2024.149485
- Primary Citation Related Structures:
8XJE, 8XJG - PubMed Abstract:
YqeY is a functionally and structurally uncharacterized protein that is ubiquitously expressed in bacteria. To gain structural insights into the function of YqeY, we determined the crystal structures of the Campylobacter jejuni and Vibrio parahaemolyticus YqeY proteins (cjYqeY and vpYqeY, respectively) and analyzed the structural and functional roles of conserved residues via a mutational study. Both cjYqeY and vpYqeY were found to adopt a two-domain structure consisting of an N-terminal four-α-helix domain and a C-terminal three-α-helix domain, with a relatively flexible interdomain orientation. The YqeY structure is unique in its linkage of the two α-helix domains although the C-terminal YqeY domain is structurally homologous to the terminal appendages of glutaminyl-tRNA synthetase and tRNA-dependent amidotransferase. We identified six conserved YqeY residues (Y67, R72, E82, Y89, P91, and G119) and evaluated their roles in protein stability via alanine mutation using a thermal shift assay. Residues Y67, R72, Y89, and P91 were shown to be required to maintain the structural integrity of YqeY. In contrast, residues E82 and G119 were not found to be essential for protein stability and are highly likely to contribute to the biological function of YqeY.
- Division of Biomedical Convergence, College of Biomedical Science, Kangwon National University, Chuncheon, 24341, Republic of Korea.
Organizational Affiliation: