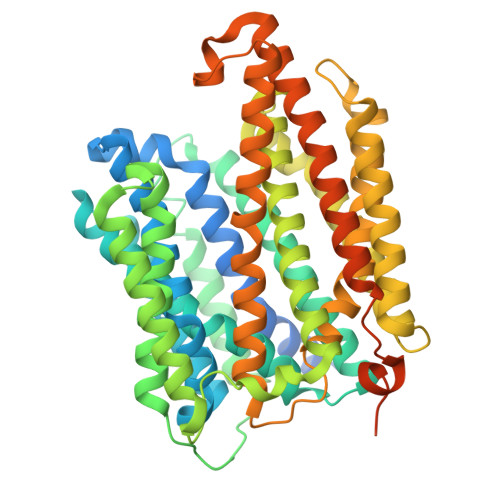
Structure and function of Mycobacterium tuberculosis EfpA as a lipid transporter and its inhibition by BRD-8000.3.
Li, D., Zhang, X., Yao, Y., Sun, X., Sun, J., Ma, X., Yuan, K., Bai, G., Pang, X., Hua, R., Guo, T., Mi, Y., Wu, L., Zhang, J., Wu, Y., Liu, Y., Wang, P., Wong, C.C.L., Chen, X.W., Xiao, H., Gao, G.F., Gao, F.(2024) Proc Natl Acad Sci U S A 121: e2412653121-e2412653121
- PubMed: 39441632 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2412653121
- Primary Citation Related Structures:
8X6X, 8YNZ - PubMed Abstract:
EfpA, the first major facilitator superfamily (MFS) protein identified in Mycobacterium tuberculosis (Mtb), is an essential efflux pump implicated in resistance to multiple drugs. EfpA-inhibitors have been developed to kill drug-tolerant Mtb. However, the biological function of EfpA has not yet been elucidated. Here, we present the cryo-EM structures of EfpA complexed with lipids or the inhibitor BRD-8000.3 at resolutions of 2.9 Å and 3.4 Å, respectively. Unexpectedly, EfpA forms an antiparallel dimer. Functional studies reveal that EfpA is a lipid transporter and BRD-8000.3 inhibits its lipid transport activity. Intriguingly, the mutation V319F, known to confer resistance to BRD-8000.3, alters the expression level and oligomeric state of EfpA. Based on our results and the observation of other antiparallel dimers in the MFS family, we propose an antiparallel-function model of EfpA. Collectively, our work provides structural and functional insights into EfpA's role in lipid transport and drug resistance, which would accelerate the development of antibiotics against this promising drug target.
- Laboratory of Protein Engineering and Vaccines, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin 300308, China.
Organizational Affiliation: