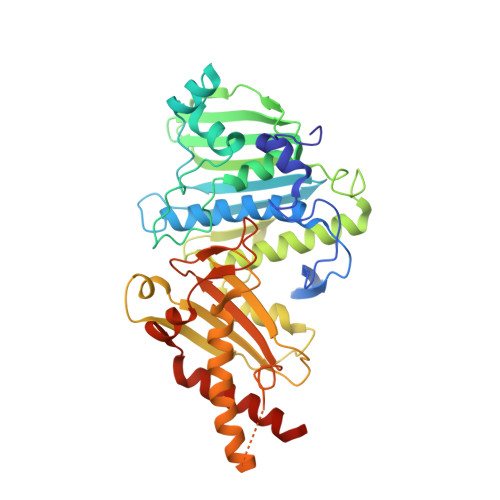
Structure-function analysis of the ATPase domain of African swine fever virus topoisomerase.
Kuang, W., Zhao, Y., Li, J., Deng, Z.(2024) mBio 15: e0308623-e0308623
- PubMed: 38411066 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.03086-23
- Primary Citation Related Structures:
8WWO - PubMed Abstract:
Type II topoisomerase utilizes the energy from ATP hydrolysis to alter DNA topology during genome replication and transcription. The ATPase domain of this enzyme is required for ATP hydrolysis and plays a crucial role in coupling DNA binding and ATP turnover with the DNA strand passage reaction. The African swine fever virus (ASFV) specifically encodes a topoisomerase II (topo II), which is critical for viral replication and an attractive target for antiviral development. Here, we present a high-resolution crystal structure of the ASFV topo II ATPase domain complexed with the substrate analog AMPPNP. Structural comparison reveals that the ASFV topo II ATPase domain shares a conserved overall structure with its homologs from eukaryotes and prokaryotes but also has three characteristic regions, including the intra-molecular interface formed by the ATP-lid and QTK loop as well as helix α9, the K-loop in the transducer domain, and the antennae-like α-helix at the ATP binding domain. Mutating the key residues within these three regions impairs or abolishes the basal and DNA-stimulated ATPase activities and reduces or eliminates the relaxation activity of the holoenzyme. Our data indicate that all three regions are functionally important for the ATPase and relaxation activities and strongly suggest that ATP hydrolysis, DNA binding, and strand passage are highly coupled and managed by the allosteric coordination of multiple domains of the type II topoisomerase. Moreover, we find a promising druggable pocket in the dimeric interface of the ASFV topo II ATPase domain, which will benefit future anti-ASFV drug development. The ATPase domain of type II topoisomerase provides energy by hydrolyzing ATP and coordinates with the DNA-binding/cleavage domain to drive and control DNA transport. The precise molecular mechanisms of how these domains respond to DNA binding and ATP hydrolysis signals and communicate with each other remain elusive. We determine the first high-resolution crystal structure of the ATPase domain of African swine fever virus (ASFV) topo II in complex with AMPPNP and biochemically investigate its function in ATPase and DNA relaxation activities. Importantly, we find that mutations at three characteristic regions of the ASFV ATPase domain produce parallel effects on the basal/DNA-stimulated ATPase and relaxation activities, implying the tight coupling of the ATP hydrolysis and strand passage process. Therefore, our data provide important implications for understanding the strand passage mechanism of the type II topoisomerase and the structural basis for developing ATPase domain-targeting antivirals against ASFV.
- Key Laboratory of Special Pathogens and Biosafety, Wuhan Institute of Virology, Center for Antiviral Research, Chinese Academy of Sciences, Wuhan, Hubei, China.
Organizational Affiliation: