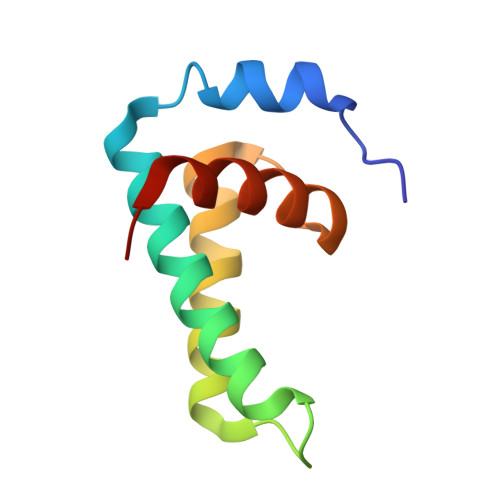
AcrIIA28 is a metalloprotein that specifically inhibits targeted-DNA loading to SpyCas9 by binding to the REC3 domain.
Kim, G.E., Park, H.H.(2024) Nucleic Acids Res 52: 6459-6471
- PubMed: 38726868 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkae357
- Primary Citation Related Structures:
8WRX - PubMed Abstract:
CRISPR-Cas systems serve as adaptive immune systems in bacteria and archaea, protecting against phages and other mobile genetic elements. However, phages and archaeal viruses have developed countermeasures, employing anti-CRISPR (Acr) proteins to counteract CRISPR-Cas systems. Despite the revolutionary impact of CRISPR-Cas systems on genome editing, concerns persist regarding potential off-target effects. Therefore, understanding the structural and molecular intricacies of diverse Acrs is crucial for elucidating the fundamental mechanisms governing CRISPR-Cas regulation. In this study, we present the structure of AcrIIA28 from Streptococcus phage Javan 128 and analyze its structural and functional features to comprehend the mechanisms involved in its inhibition of Cas9. Our current study reveals that AcrIIA28 is a metalloprotein that contains Zn2+ and abolishes the cleavage activity of Cas9 only from Streptococcus pyrogen (SpyCas9) by directly interacting with the REC3 domain of SpyCas9. Furthermore, we demonstrate that the AcrIIA28 interaction prevents the target DNA from being loaded onto Cas9. These findings indicate the molecular mechanisms underlying AcrIIA28-mediated Cas9 inhibition and provide valuable insights into the ongoing evolutionary battle between bacteria and phages.
- College of Pharmacy, Chung-Ang University, Seoul 06974, Republic of Korea.
Organizational Affiliation: