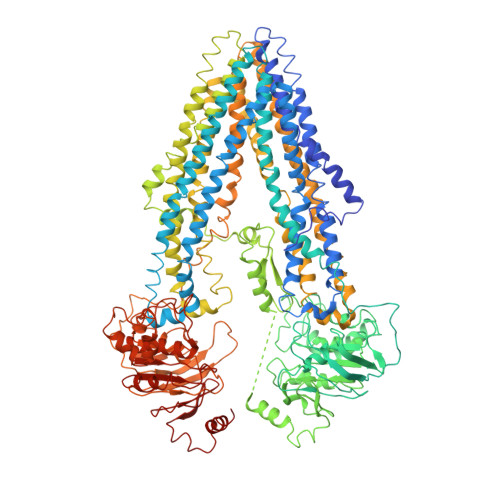
Structure and function of the Arabidopsis ABC transporter ABCB19 in brassinosteroid export.
Ying, W., Wang, Y., Wei, H., Luo, Y., Ma, Q., Zhu, H., Janssens, H., Vukasinovic, N., Kvasnica, M., Winne, J.M., Gao, Y., Tan, S., Friml, J., Liu, X., Russinova, E., Sun, L.(2024) Science 383: eadj4591-eadj4591
- PubMed: 38513023 Search on PubMed
- DOI: https://doi.org/10.1126/science.adj4591
- Primary Citation Related Structures:
8WOI, 8WOM, 8WOO, 8WP0 - PubMed Abstract:
Brassinosteroids are steroidal phytohormones that regulate plant development and physiology, including adaptation to environmental stresses. Brassinosteroids are synthesized in the cell interior but bind receptors at the cell surface, necessitating a yet to be identified export mechanism. Here, we show that a member of the ATP-binding cassette (ABC) transporter superfamily, ABCB19, functions as a brassinosteroid exporter. We present its structure in both the substrate-unbound and the brassinosteroid-bound states. Bioactive brassinosteroids are potent activators of ABCB19 ATP hydrolysis activity, and transport assays showed that ABCB19 transports brassinosteroids. In Arabidopsis thaliana , ABCB19 and its close homolog, ABCB1, positively regulate brassinosteroid responses. Our results uncover an elusive export mechanism for bioactive brassinosteroids that is tightly coordinated with brassinosteroid signaling.
- Department of Neurology of The First Affiliated Hospital of USTC, MOE Key Laboratory for Membraneless Organelles and Cellular Dynamics, Hefei National Research Center for Physical Sciences at the Microscale, Biomedical Sciences and Health Laboratory of Anhui Province, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei 230027, China.
Organizational Affiliation: