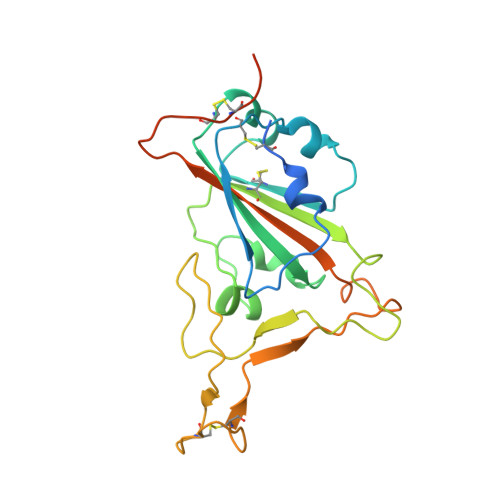
Structural basis of increased binding affinities of spikes from SARS-CoV-2 Omicron variants to rabbit and hare ACE2s reveals the expanding host tendency.
Shi, K., Li, L., Luo, C., Xu, Z., Huang, B., Ma, S., Liu, K., Yu, G., Gao, G.F.(2024) mBio 15: e0298823-e0298823
- PubMed: 38112468 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.02988-23
- Primary Citation Related Structures:
8WOX, 8WOY, 8WOZ - PubMed Abstract:
The potential host range of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has been expanding alongside its evolution during the pandemic, with rabbits and hares being considered important potential hosts, supported by a report of rabbit sero-prevalence in nature. We measured the binding affinities of rabbit and hare angiotensin-converting enzyme 2 (ACE2) with receptor-binding domains (RBDs) from SARS-CoV, SARS-CoV-2, and its variants and found that rabbit and hare ACE2s had broad variant tropism, with significantly enhanced affinities to Omicron BA.4/5 and its subsequent-emerged sub-variants (>10 fold). The structures of rabbit ACE2 complexed with either SARS-CoV-2 prototype (PT) or Omicron BA.4/5 spike (S) proteins were determined, thereby unveiling the importance of rabbit ACE2 Q34 in RBD-interaction and elucidating the molecular basis of the enhanced binding with Omicron BA.4/5 RBD. These results address the highly enhanced risk of rabbits infecting SARS-CoV-2 Omicron sub-variants and the importance of constant surveillance.IMPORTANCEThe severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic has swept the globe and caused immense health and economic damage. SARS-CoV-2 has demonstrated a broad host range, indicating a high risk of interspecies transmission and adaptive mutation. Therefore, constant monitoring for potential hosts is of immense importance. In this study, we found that Omicron BA.4/5 and subsequent-emerged sub-variants exhibited enhanced binding to both rabbit and hare angiotensin-converting enzyme 2 (ACE2), and we elucidated the structural mechanism of their recognition. From the structure, we found that Q34, a unique residue of rabbit ACE2 compared to other ACE2 orthologs, plays an important role in ACE2 recognition. These results address the probability of rabbits/hares being potential hosts of SARS-CoV-2 and broaden our knowledge regarding the molecular mechanism of SARS-CoV-2 interspecies transmission.
- Hubei Provincial Key Laboratory for Protection and Application of Special Plants in Wuling Area of China, College of Life Sciences, South-Central Minzu University, Wuhan, China.
Organizational Affiliation: